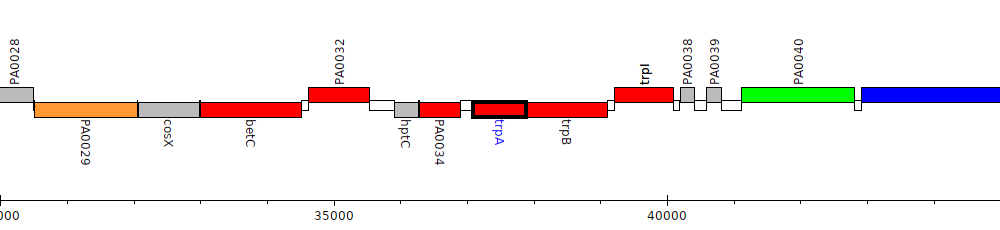
Pseudomonas aeruginosa PAO1, PA0035 (trpA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0000162 | tryptophan biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3127651 | Reviewed by curator |
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3127651 | Reviewed by curator |
| Molecular Function | GO:0004834 | tryptophan synthase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3127651 | Reviewed by curator |
| Biological Process | GO:0000162 | tryptophan biosynthetic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
3127651 | Reviewed by curator |
| Biological Process | GO:0000162 | tryptophan biosynthetic process | Non-traceable Author Statement | ECO:0000303 non-traceable author statement used in manual assertion |
3127651 | Reviewed by curator |
| Molecular Function | GO:0004834 | tryptophan synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00290
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006568 | tryptophan metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00290
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Phenylalanine, tyrosine and tryptophan biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | TRPSYN-PWY | L-tryptophan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | ALL-CHORISMATE-PWY | superpathway of chorismate metabolism | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | COMPLETE-ARO-PWY | superpathway of aromatic amino acid biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00290 | Tryptophan synthase alpha chain | IPR002028 | Tryptophan synthase, alpha chain | 8 | 263 | 3.2E-106 |
| Hamap | MF_00131 | Tryptophan synthase alpha chain [trpA]. | IPR002028 | Tryptophan synthase, alpha chain | 14 | 263 | 38.09428 |
| SUPERFAMILY | SSF51366 | Ribulose-phoshate binding barrel | IPR011060 | Ribulose-phosphate binding barrel | 1 | 248 | 1.51E-92 |
| NCBIfam | TIGR00262 | JCVI: tryptophan synthase subunit alpha | IPR002028 | Tryptophan synthase, alpha chain | 8 | 259 | 3.2E-81 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 1 | 268 | 1.9E-104 |
| PANTHER | PTHR43406 | TRYPTOPHAN SYNTHASE, ALPHA CHAIN | IPR002028 | Tryptophan synthase, alpha chain | 3 | 262 | 8.2E-85 |
| CDD | cd04724 | Tryptophan_synthase_alpha | IPR002028 | Tryptophan synthase, alpha chain | 18 | 260 | 2.07603E-121 |
| FunFam | G3DSA:3.20.20.70:FF:000037 | Tryptophan synthase alpha chain | - | - | 1 | 266 | 4.4E-101 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.