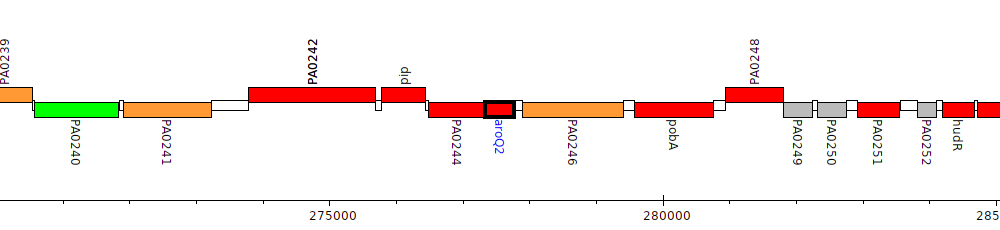
Pseudomonas aeruginosa PAO1, PA0245 (aroQ2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006571 | tyrosine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004334 | fumarylacetoacetase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0000162 | tryptophan biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0009094 | L-phenylalanine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0003855 | 3-dehydroquinate dehydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00466
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-6163 | chorismate biosynthesis from 3-dehydroquinate | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | ALL-CHORISMATE-PWY | superpathway of chorismate metabolism | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | ARO-PWY | chorismate biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Phenylalanine, tyrosine and tryptophan biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | COMPLETE-ARO-PWY | superpathway of aromatic amino acid biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00466 | DHQase_II | IPR001874 | Dehydroquinase, class II | 4 | 143 | 2.79938E-91 |
| PIRSF | PIRSF001399 | DHQase_2 | IPR001874 | Dehydroquinase, class II | 1 | 147 | 3.8E-63 |
| Hamap | MF_00169 | 3-dehydroquinate dehydratase [aroQ]. | IPR001874 | Dehydroquinase, class II | 1 | 148 | 73.130402 |
| SUPERFAMILY | SSF52304 | Type II 3-dehydroquinate dehydratase | IPR036441 | Dehydroquinase, class II superfamily | 4 | 145 | 1.27E-62 |
| Gene3D | G3DSA:3.40.50.9100 | Dehydroquinase, class II | IPR036441 | Dehydroquinase, class II superfamily | 1 | 148 | 1.5E-69 |
| PANTHER | PTHR21272 | CATABOLIC 3-DEHYDROQUINASE | IPR001874 | Dehydroquinase, class II | 1 | 146 | 3.8E-63 |
| Pfam | PF01220 | Dehydroquinase class II | IPR001874 | Dehydroquinase, class II | 5 | 140 | 2.1E-61 |
| NCBIfam | TIGR01088 | JCVI: 3-dehydroquinate dehydratase, type II | IPR001874 | Dehydroquinase, class II | 5 | 142 | 2.3E-61 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.