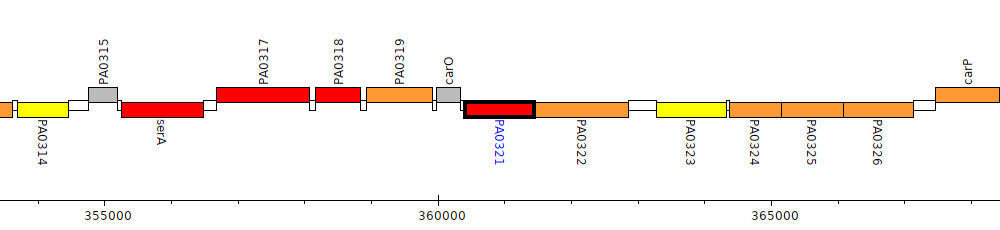
Pseudomonas aeruginosa PAO1, PA0321
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006595 | polyamine metabolic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
26956223 | Reviewed by curator |
| Molecular Function | GO:0016787 | hydrolase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
26956223 | Reviewed by curator |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00850 | Histone deacetylase domain | IPR023801 | Histone deacetylase domain | 28 | 336 | 7.6E-63 |
| PANTHER | PTHR48252 | HISTONE DEACETYLASE 2-RELATED | - | - | 24 | 328 | 3.2E-50 |
| Gene3D | G3DSA:3.40.800.20 | Histone deacetylase domain | IPR037138 | Histone deacetylase domain superfamily | 1 | 343 | 1.8E-123 |
| PRINTS | PR01270 | Histone deacetylase superfamily signature | IPR000286 | Histone deacetylase family | 188 | 203 | 1.0E-12 |
| SUPERFAMILY | SSF52768 | Arginase/deacetylase | IPR023696 | Ureohydrolase domain superfamily | 2 | 341 | 1.15E-85 |
| CDD | cd10001 | HDAC_classII_APAH | - | - | 2 | 340 | 0.0 |
| PRINTS | PR01270 | Histone deacetylase superfamily signature | IPR000286 | Histone deacetylase family | 274 | 284 | 1.0E-12 |
| PRINTS | PR01270 | Histone deacetylase superfamily signature | IPR000286 | Histone deacetylase family | 155 | 178 | 1.0E-12 |
| FunFam | G3DSA:3.40.800.20:FF:000032 | Acetylpolyamine amidohydrolase 2 | - | - | 1 | 343 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.