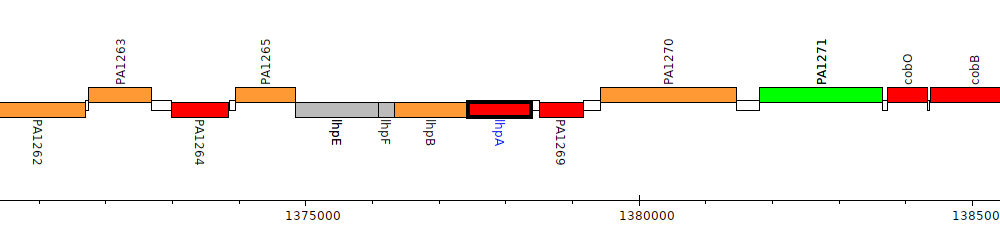
Pseudomonas aeruginosa PAO1, PA1268 (lhpA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0047580 | 4-hydroxyproline epimerase activity | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
27145750 | Reviewed by curator |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | PWY-5159 | trans-4-hydroxy-L-proline degradation II | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00330 | Arginine and proline metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF54506 | Diaminopimelate epimerase-like | - | - | 2 | 312 | 5.34E-100 |
| Pfam | PF05544 | Proline racemase | IPR008794 | Proline racemase family | 6 | 312 | 2.6E-111 |
| Gene3D | G3DSA:3.10.310.10 | Diaminopimelate Epimerase; Chain A, domain 1 | - | - | 4 | 311 | 0.0 |
| Gene3D | G3DSA:3.10.310.10 | Diaminopimelate Epimerase; Chain A, domain 1 | - | - | 134 | 290 | 0.0 |
| PANTHER | PTHR33442 | TRANS-3-HYDROXY-L-PROLINE DEHYDRATASE | IPR008794 | Proline racemase family | 3 | 312 | 4.8E-80 |
| SFLD | SFLDS00028 | Proline Racemase | IPR008794 | Proline racemase family | 2 | 313 | 0.0 |
| FunFam | G3DSA:3.10.310.10:FF:000012 | 4-hydroxyproline 2-epimerase | - | - | 134 | 290 | 2.6E-78 |
| PIRSF | PIRSF029792 | Pro_racemase | IPR008794 | Proline racemase family | 1 | 313 | 1.7E-94 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.