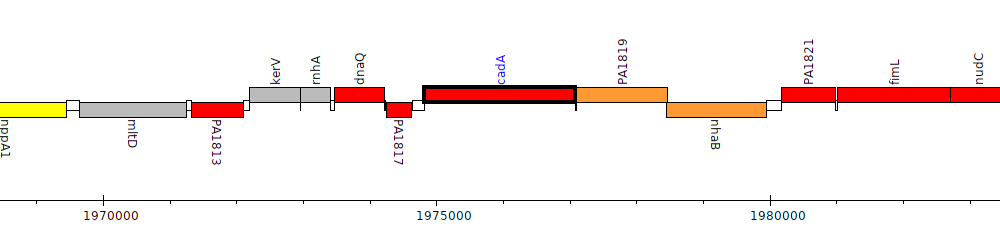
Pseudomonas aeruginosa PAO1, PA1818 (cadA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009393
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01276
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009393
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016831 | carboxy-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF009393
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glutamate metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00330 | Arginine and proline metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Lysine degradation |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Arginine and proline metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Urea cycle and metabolism of amino groups |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53383 | PLP-dependent transferases | IPR015424 | Pyridoxal phosphate-dependent transferase | 149 | 599 | 1.21E-112 |
| Pfam | PF01276 | Orn/Lys/Arg decarboxylase, major domain | IPR000310 | Orn/Lys/Arg decarboxylase, major domain | 149 | 582 | 0.0 |
| CDD | cd00615 | Orn_deC_like | IPR000310 | Orn/Lys/Arg decarboxylase, major domain | 150 | 469 | 7.24844E-123 |
| PIRSF | PIRSF009393 | Orn_decarb | IPR011193 | Ornithine/lysine/arginine decarboxylase | 3 | 751 | 0.0 |
| Pfam | PF03711 | Orn/Lys/Arg decarboxylase, C-terminal domain | IPR008286 | Orn/Lys/Arg decarboxylase, C-terminal | 607 | 736 | 7.6E-49 |
| FunFam | G3DSA:3.40.640.10:FF:000008 | Lysine decarboxylase, inducible | - | - | 149 | 454 | 6.2E-125 |
| PANTHER | PTHR45229 | CONSTITUTIVE ORNITHINE DECARBOXYLASE | IPR011193 | Ornithine/lysine/arginine decarboxylase | 10 | 744 | 0.0 |
| Pfam | PF03709 | Orn/Lys/Arg decarboxylase, N-terminal domain | IPR005308 | Orn/Lys/Arg decarboxylase, N-terminal | 22 | 143 | 6.0E-24 |
| Gene3D | G3DSA:3.90.1150.10 | Aspartate Aminotransferase, domain 1 | IPR015422 | Pyridoxal phosphate-dependent transferase, small domain | 455 | 627 | 1.4E-64 |
| Gene3D | G3DSA:3.90.100.10 | - | - | - | 628 | 751 | 1.7E-43 |
| Gene3D | G3DSA:3.40.640.10 | - | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 149 | 454 | 2.3E-100 |
| Gene3D | G3DSA:3.40.50.2300 | - | - | - | 8 | 148 | 2.7E-31 |
| SUPERFAMILY | SSF55904 | Ornithine decarboxylase C-terminal domain | IPR036633 | Orn/Lys/Arg decarboxylase, C-terminal domain superfamily | 602 | 751 | 1.24E-43 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.