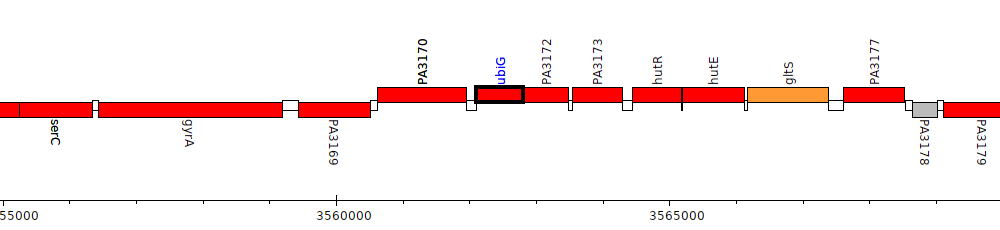
Pseudomonas aeruginosa PAO1, PA3171 (ubiG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0018467 | formaldehyde dehydrogenase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0051188 | obsolete cofactor biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53335
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008425 | 2-polyprenyl-6-methoxy-1,4-benzoquinone methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00472
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006744 | ubiquinone biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00472
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | ubiquinol-8 biosynthesis (prokaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | ALL-CHORISMATE-PWY | superpathway of chorismate metabolism | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| MetaCyc | ubiquinol-9 biosynthesis (prokaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pae00130 | Ubiquinone and other terpenoid-quinone biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | ubiquinol-9 biosynthesis (eukaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ubiquinol-10 biosynthesis (eukaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ubiquinol-6 bypass biosynthesis (eukaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCyc | PWY-6708 | ubiquinol-8 biosynthesis (prokaryotic) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| MetaCyc | ubiquinol-10 biosynthesis (prokaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ubiquinol-8 biosynthesis (eukaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ubiquinol-7 biosynthesis (prokaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Ubiquinone biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | ubiquinol-7 biosynthesis (eukaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ubiquinol-6 biosynthesis from 4-aminobenzoate (yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF13489 | Methyltransferase domain | - | - | 47 | 199 | 2.3E-27 |
| SUPERFAMILY | SSF53335 | S-adenosyl-L-methionine-dependent methyltransferases | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 12 | 226 | 8.83E-52 |
| Gene3D | G3DSA:3.40.50.150 | Vaccinia Virus protein VP39 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 8 | 231 | 7.3E-80 |
| PANTHER | PTHR43464 | METHYLTRANSFERASE | - | - | 2 | 192 | 5.1E-37 |
| CDD | cd02440 | AdoMet_MTases | - | - | 50 | 149 | 7.35848E-16 |
| NCBIfam | TIGR01983 | JCVI: bifunctional 2-polyprenyl-6-hydroxyphenol methylase/3-demethylubiquinol 3-O-methyltransferase UbiG | IPR010233 | Ubiquinone biosynthesis O-methyltransferase | 8 | 225 | 7.2E-87 |
| Hamap | MF_00472 | Ubiquinone biosynthesis O-methyltransferase [ubiG]. | IPR010233 | Ubiquinone biosynthesis O-methyltransferase | 8 | 231 | 37.630657 |
| FunFam | G3DSA:3.40.50.150:FF:000028 | Ubiquinone biosynthesis O-methyltransferase | - | - | 8 | 231 | 6.9E-105 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.