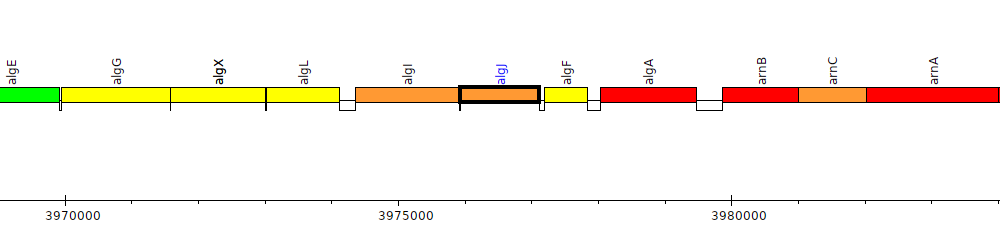
Pseudomonas aeruginosa PAO1, PA3549 (algJ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0042121 | alginic acid biosynthetic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8636017 | Reviewed by curator |
| Biological Process | GO:0051979 | alginic acid acetylation | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8636017 | Reviewed by curator |
| Biological Process | GO:0051979 | alginic acid acetylation | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
12003941 | Reviewed by curator |
| Biological Process | GO:0050896 | response to stimulus | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8636017 | Reviewed by curator |
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0042121 | alginic acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd14442
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Alginate biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PWY-6082 | alginate biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 259 | 286 | - |
| Pfam | PF16822 | SGNH hydrolase-like domain, acetyltransferase AlgX | IPR031811 | AlgX/AlgJ, SGNH hydrolase-like domain | 85 | 355 | 2.0E-96 |
| CDD | cd14442 | AlgJ_like | IPR034657 | Alginate O-acetyltranferase AlgJ | 47 | 374 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.