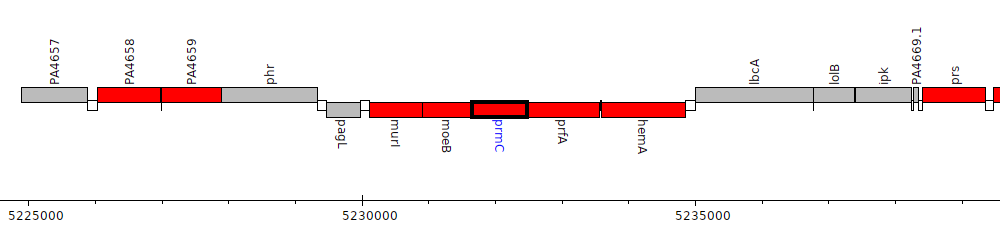
Pseudomonas aeruginosa PAO1, PA4664 (prmC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA14_61680
|
ECO:0000250 sequence similarity evidence used in manual assertion |
23278968 | Reviewed by curator |
| Biological Process | GO:0006479 | protein methylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00536
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008276 | protein methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00536
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53335
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03534
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | CODH-PWY | reductive acetyl coenzyme A pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Porphyrin and chlorophyll metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.150 | Vaccinia Virus protein VP39 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 84 | 275 | 5.9E-60 |
| Coils | Coil | Coil | - | - | 138 | 158 | - |
| SUPERFAMILY | SSF53335 | S-adenosyl-L-methionine-dependent methyltransferases | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 12 | 272 | 1.13E-52 |
| Pfam | PF05175 | Methyltransferase small domain | IPR007848 | Methyltransferase small domain | 94 | 185 | 1.4E-15 |
| NCBIfam | TIGR03534 | JCVI: peptide chain release factor N(5)-glutamine methyltransferase | IPR019874 | Protein-(glutamine-N5) methyltransferase, release factor-specific | 22 | 270 | 1.1E-88 |
| Gene3D | G3DSA:1.10.8.10 | - | - | - | 7 | 82 | 1.9E-21 |
| Hamap | MF_02126 | Release factor glutamine methyltransferase [prmC]. | IPR019874 | Protein-(glutamine-N5) methyltransferase, release factor-specific | 3 | 274 | 46.551399 |
| Pfam | PF17827 | PrmC N-terminal domain | IPR040758 | Release factor glutamine methyltransferase, N-terminal domain | 16 | 72 | 2.1E-15 |
| PANTHER | PTHR18895 | HEMK METHYLTRANSFERASE | - | - | 15 | 272 | 8.3E-68 |
| NCBIfam | TIGR00536 | JCVI: HemK family protein methyltransferase | IPR004556 | Methyltransferase HemK-like | 18 | 276 | 2.8E-77 |
| CDD | cd02440 | AdoMet_MTases | - | - | 112 | 236 | 2.10102E-11 |
| FunFam | G3DSA:1.10.8.10:FF:000141 | Release factor glutamine methyltransferase | - | - | 7 | 82 | 1.3E-49 |
| FunFam | G3DSA:3.40.50.150:FF:000053 | Release factor glutamine methyltransferase | - | - | 84 | 275 | 1.3E-83 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.