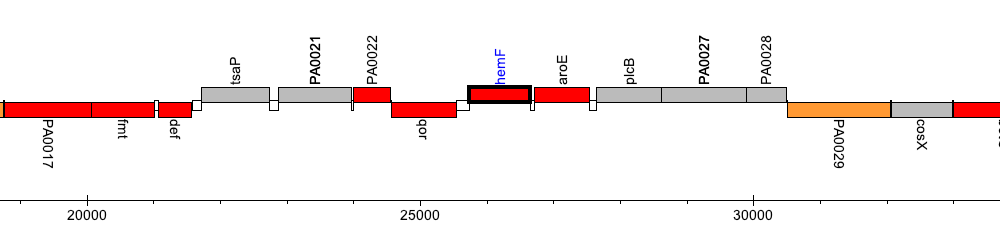
Pseudomonas aeruginosa PAO1, PA0024 (hemF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004109 | coproporphyrinogen oxidase activity | Inferred from Genetic Interaction | ECO:0000316 genetic interaction evidence used in manual assertion |
9767567 | Reviewed by curator |
| Biological Process | GO:0051188 | obsolete cofactor biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0015994 | chlorophyll metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004109 | coproporphyrinogen oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1500.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006779 | porphyrin-containing compound biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1500.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Porphyrin and chlorophyll metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00860 | Porphyrin and chlorophyll metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 161 | 190 | 5.1E-88 |
| Pfam | PF01218 | Coproporphyrinogen III oxidase | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 8 | 300 | 0.0 |
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 30 | 46 | 5.1E-88 |
| PIRSF | PIRSF000166 | Coproporphyri_ox | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 1 | 302 | 4.5E-128 |
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 233 | 260 | 5.1E-88 |
| Hamap | MF_00333 | Oxygen-dependent coproporphyrinogen-III oxidase [hemF]. | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 2 | 276 | 56.336945 |
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 261 | 287 | 5.1E-88 |
| PANTHER | PTHR10755 | COPROPORPHYRINOGEN III OXIDASE, MITOCHONDRIAL | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 7 | 301 | 1.8E-126 |
| SUPERFAMILY | SSF102886 | Coproporphyrinogen III oxidase | IPR036406 | Oxygen-dependent coproporphyrinogen III oxidase superfamily | 3 | 301 | 0.0 |
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 124 | 146 | 5.1E-88 |
| FunFam | G3DSA:3.40.1500.10:FF:000001 | Oxygen-dependent coproporphyrinogen-III oxidase | - | - | 1 | 301 | 0.0 |
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 47 | 68 | 5.1E-88 |
| Gene3D | G3DSA:3.40.1500.10 | Coproporphyrinogen III oxidase, aerobic | IPR036406 | Oxygen-dependent coproporphyrinogen III oxidase superfamily | 1 | 304 | 0.0 |
| PRINTS | PR00073 | Coprogen oxidase signature | IPR001260 | Coproporphyrinogen III oxidase, aerobic | 89 | 113 | 5.1E-88 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.