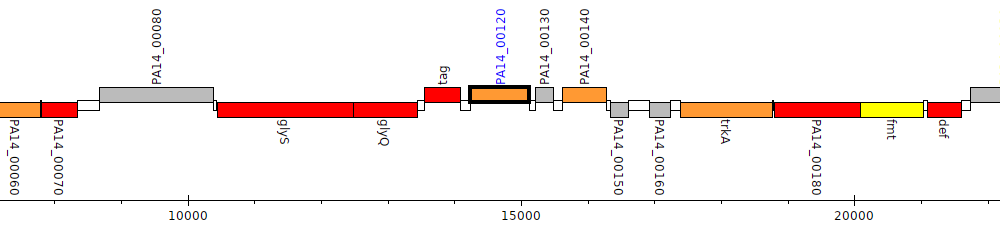
Pseudomonas aeruginosa UCBPP-PA14, PA14_00120
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0071236 | cellular response to antibiotic |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0011
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
22146126 | |
| Biological Process | GO:0034605 | cellular response to heat |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0011
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
22146126 | |
| Biological Process | GO:0071470 | cellular response to osmotic stress |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0011
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
22146126 | |
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:cd07984
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd07984
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00540 | Lipopolysaccharide biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd07984 | LPLAT_LABLAT-like | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 97 | 285 | 6.36917E-43 |
| Pfam | PF03279 | Bacterial lipid A biosynthesis acyltransferase | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 8 | 285 | 1.5E-44 |
| PANTHER | PTHR30606 | LIPID A BIOSYNTHESIS LAUROYL ACYLTRANSFERASE | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 5 | 294 | 4.9E-51 |
| PIRSF | PIRSF026649 | MsbB | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 1 | 295 | 1.0E-89 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.