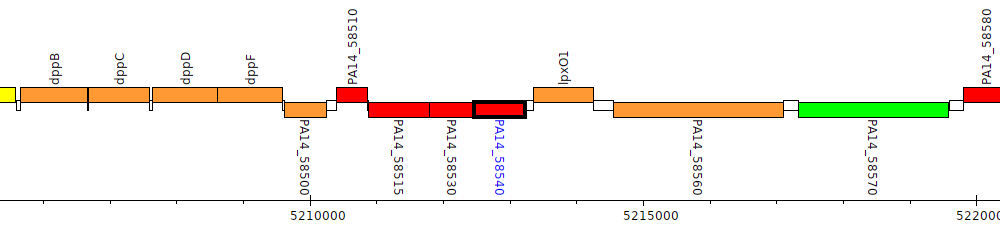
Pseudomonas aeruginosa UCBPP-PA14, PA14_58540
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF88713
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | γ-glutamyl cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 5-oxo-L-proline metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd10787 | LamB_YcsF_like | - | - | 8 | 248 | 1.66909E-130 |
| Pfam | PF03746 | LamB/YcsF family | IPR005501 | LamB/YcsF/PxpA-like | 8 | 249 | 1.1E-92 |
| Gene3D | G3DSA:3.20.20.370 | Glycoside hydrolase/deacetylase | - | - | 1 | 251 | 2.3E-105 |
| Hamap | MF_00691 | 5-oxoprolinase subunit A [pxpA]. | IPR005501 | LamB/YcsF/PxpA-like | 7 | 251 | 42.711975 |
| PANTHER | PTHR30292 | UNCHARACTERIZED PROTEIN YBGL-RELATED | IPR005501 | LamB/YcsF/PxpA-like | 6 | 249 | 2.6E-95 |
| SUPERFAMILY | SSF88713 | Glycoside hydrolase/deacetylase | IPR011330 | Glycoside hydrolase/deacetylase, beta/alpha-barrel | 7 | 249 | 3.93E-90 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.