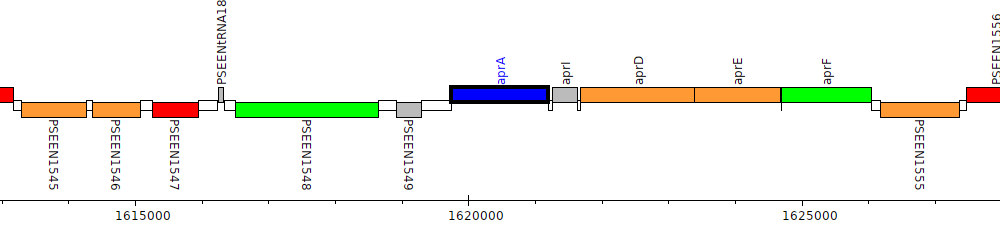
Pseudomonas entomophila L48, PSEEN1550 (aprA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0042783 | evasion of host immune response | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
16789834 | Reviewed by curator |
| Molecular Function | GO:0008233 | peptidase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
16789834 | Reviewed by curator |
| Cellular Component | GO:0005576 | extracellular region | Inferred from Direct Assay | ECO:0000333 sodium dodecyl sulfate polyacrylamide gel electrophoresis evidence |
16789834 | Reviewed by curator |
| Cellular Component | GO:0005615 | extracellular space |
Inferred from Sequence Model
Term mapped from: InterPro:PF08548
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005509 | calcium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00353
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008237 | metallopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.390.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pen01503 | Cationic antimicrobial peptide (CAMP) resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF08548 | Peptidase M10 serralysin C terminal | IPR013858 | Peptidase M10 serralysin, C-terminal | 263 | 485 | 6.8E-79 |
| PRINTS | PR00313 | NodO calcium binding signature | - | - | 345 | 356 | 2.9E-12 |
| Pfam | PF13583 | Metallo-peptidase family M12B Reprolysin-like | - | - | 175 | 240 | 2.8E-6 |
| PRINTS | PR00313 | NodO calcium binding signature | - | - | 375 | 383 | 2.9E-12 |
| Gene3D | G3DSA:3.40.390.10 | Collagenase (Catalytic Domain) | IPR024079 | Metallopeptidase, catalytic domain superfamily | 37 | 265 | 0.0 |
| FunFam | G3DSA:3.40.390.10:FF:000046 | Serralysin | - | - | 36 | 265 | 4.8E-116 |
| PRINTS | PR00313 | NodO calcium binding signature | - | - | 366 | 374 | 2.9E-12 |
| SUPERFAMILY | SSF55486 | Metalloproteases ("zincins"), catalytic domain | - | - | 27 | 261 | 3.42E-66 |
| PRINTS | PR00313 | NodO calcium binding signature | - | - | 384 | 393 | 2.9E-12 |
| NCBIfam | NF035945 | NCBIFAM: serralysin family metalloprotease | - | - | 24 | 485 | 0.0 |
| PIRSF | PIRSF001205 | Serralysin | IPR016294 | Peptidase M10B | 5 | 485 | 0.0 |
| PANTHER | PTHR10201 | MATRIX METALLOPROTEINASE | - | - | 74 | 285 | 4.9E-9 |
| Pfam | PF00353 | RTX calcium-binding nonapeptide repeat (4 copies) | IPR001343 | RTX calcium-binding nonapeptide repeat | 356 | 390 | 6.6E-10 |
| SMART | SM00235 | col_5 | IPR006026 | Peptidase, metallopeptidase | 79 | 243 | 1.4E-19 |
| SUPERFAMILY | SSF51120 | beta-Roll | IPR011049 | Serralysin-like metalloprotease, C-terminal | 263 | 484 | 3.79E-49 |
| CDD | cd04277 | ZnMc_serralysin_like | IPR034033 | Serralysin-like metallopeptidase domain | 72 | 262 | 7.04096E-43 |
| FunFam | G3DSA:2.150.10.10:FF:000001 | Serralysin | - | - | 260 | 482 | 1.3E-116 |
| Gene3D | G3DSA:2.150.10.10 | - | IPR011049 | Serralysin-like metalloprotease, C-terminal | 31 | 482 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.