Pseudomonas aeruginosa PAO1, PA3785
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
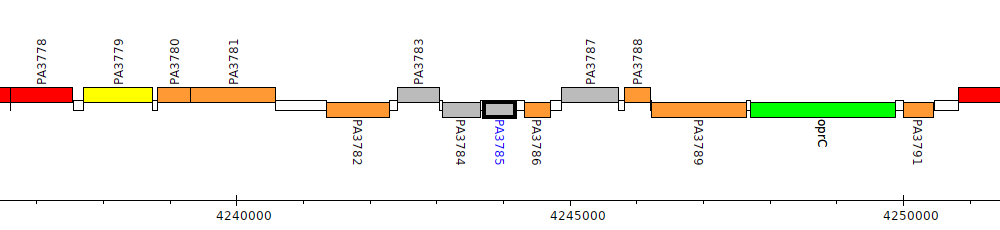
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA3785
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 4243709 - 4244185 (- strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Comment | Structural similarity to 2K6W: copper protein from Thermus thermophilus (root-mean-square deviation: 1.5 angstroms) |
Cross-References
| RefSeq | NP_252474.1 |
| GI | 15598980 |
| Affymetrix | PA3785_at |
| DNASU | PaCD00007649 |
| Entrez | 879117 |
| GenBank | AAG07172.1 |
| INSDC | AAG07172.1 |
| NCBI Locus Tag | PA3785 |
| protein_id(GenBank) | gb|AAG07172.1|AE004797_7|gnl|PseudoCAP|PA3785 |
| TIGR | NTL03PA03785 |
| UniParc | UPI00000C5B10 |
| UniProtKB Acc | Q9HXL0 |
| UniProtKB ID | Q9HXL0_PSEAE |
| UniRef100 | UniRef100_W0YXG0 |
| UniRef50 | UniRef50_Q87VQ4 |
| UniRef90 | UniRef90_A6V0X9 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
putative copper binding protein
|
| Synonyms | |
| Evidence for Translation |
Detected by MALDI-TOF/TOF (PMID:19333994).
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -1.13 |
| Kyte-Doolittle Hydrophobicity Value | -0.216 |
| Molecular Weight (kDa) | 17.0 |
| Isoelectric Point (pI) | 6.80 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 202 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001881 (481 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA3785
Search term: putative copper binding protein
|
Human Homologs
References
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|
|
Uncovering the structure and function of Pseudomonas aeruginosa periplasmic proteins by an in silico approach.
Caprari S, Brandi V, Pasquadibisceglie A, Polticelli F
J Biomol Struct Dyn 2020 Sep;38(15):4508-4520
PubMed ID: 31631799
|