Pseudomonas aeruginosa PAO1, PA4723
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
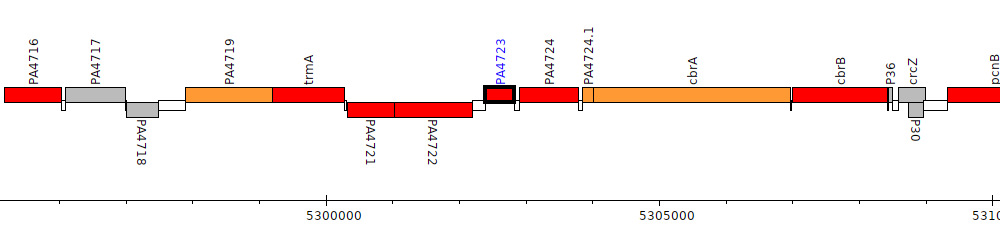
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA4723
|
| Name |
Synonym: dksA1 |
| Replicon | chromosome |
| Genomic location | 5302386 - 5302832 (+ strand) |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
| Comment | DksA1 is required for H2O2 tolerance in both planktonic and biofilm growing cells, and protects against macrophage-mediated killing |
Cross-References
| RefSeq | NP_253411.1 |
| GI | 15599917 |
| Affymetrix | PA4723_dksA_at |
| Entrez | 881594 |
| GenBank | AAG08109.1 |
| INSDC | AAG08109.1 |
| NCBI Locus Tag | PA4723 |
| protein_id(GenBank) | gb|AAG08109.1|AE004886_1|gnl|PseudoCAP|PA4723 |
| TIGR | NTL03PA04724 |
| UniParc | UPI00000C4ED2 |
| UniProtKB Acc | G3XD14 |
| UniProtKB ID | G3XD14_PSEAE |
| UniRef100 | UniRef100_W0YXF7 |
| UniRef50 | UniRef50_P43758 |
| UniRef90 | UniRef90_S6BNJ9 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
suppressor protein DksA
|
| Synonyms | |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
Detected by MALDI-TOF/TOF (PMID:19333994).
Detected by two-dimensional gel electrophoresis, mass spectral analysis and protein mass mapping (PMID:21863142).
Verified using two-dimensional gel electrophoresis, MALDI-TOF MS and peptide mass mapping(pI:5.0, molecular weight:17.3, PMID:21674802)
|
| Charge (pH 7) | -7.40 |
| Kyte-Doolittle Hydrophobicity Value | -0.959 |
| Molecular Weight (kDa) | 17.3 |
| Isoelectric Point (pI) | 4.77 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 331 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG004369 (536 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA4723
Search term: suppressor protein DksA
|
Human Homologs
References
|
Inhibition of quorum sensing by a Pseudomonas aeruginosa dksA homologue.
Branny P, Pearson JP, Pesci EC, Köhler T, Iglewski BH, Van Delden C
J Bacteriol 2001 Mar;183(5):1531-9
PubMed ID: 11160083
|
|
Nucleotide sequence and characterization of the sfs1 gene: sfs1 is involved in CRP*-dependent mal gene expression in Escherichia coli.
Kawamukai M, Utsumi R, Takeda K, Higashi A, Matsuda H, Choi YL, Komano T
J Bacteriol 1991 Apr;173(8):2644-8
PubMed ID: 2013578
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|
|
Comparisons of Two Proteomic Analyses of Non-Mucoid and Mucoid Pseudomonas aeruginosa Clinical Isolates from a Cystic Fibrosis Patient.
Rao J, Damron FH, Basler M, Digiandomenico A, Sherman NE, Fox JW, Mekalanos JJ, Goldberg JB
Front Microbiol 2011;2:162
PubMed ID: 21863142
|
|
Proteomics of the oxidative stress response induced by hydrogen peroxide and paraquat reveals a novel AhpC-like protein in Pseudomonas aeruginosa.
Hare NJ, Scott NE, Shin EH, Connolly AM, Larsen MR, Palmisano G, Cordwell SJ
Proteomics 2011 Aug;11(15):3056-69
PubMed ID: 21674802
|
|
Role of a Zn-independent DksA in Zn homeostasis and stringent response.
Blaby-Haas CE, Furman R, Rodionov DA, Artsimovitch I, de Crécy-Lagard V
Mol Microbiol 2011 Feb;79(3):700-15
PubMed ID: 21255113
|
|
The Pseudomonas aeruginosa DksA1 protein is involved in H(2)O(2) tolerance and within-macrophages survival and can be replaced by DksA2.
Fortuna A, Collalto D, Schiaffi V, Pastore V, Visca P, Ascenzioni F, Rampioni G, Leoni L
Sci Rep 2022 Jun 21;12(1):10404
PubMed ID: 35729352
|