Pseudomonas aeruginosa PAO1, PA5134 (ctpA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
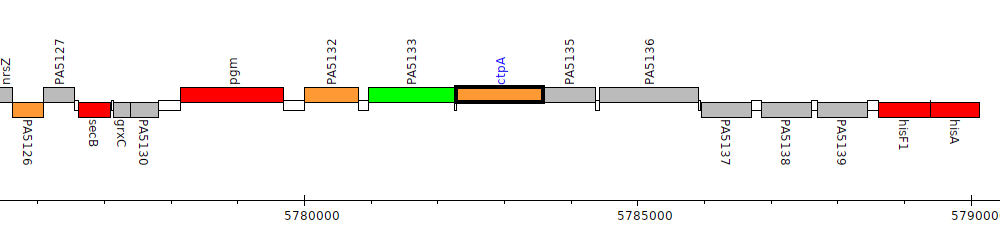
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA5134
|
| Name |
ctpA
Synonym: ctpA |
| Replicon | chromosome |
| Genomic location | 5782278 - 5783588 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
| Comment | Structural similarity to 4C2E: carboxyprotease from Tetradesmus obliquus (root-mean-square deviation: 1.6 angstroms) |
Cross-References
| RefSeq | NP_253821.1 |
| GI | 15600327 |
| Affymetrix | PA5134_at |
| Entrez | 878610 |
| GenBank | AAG08519.1 |
| INSDC | AAG08519.1 |
| NCBI Locus Tag | PA5134 |
| protein_id(GenBank) | gb|AAG08519.1|AE004926_9|gnl|PseudoCAP|PA5134 |
| TIGR | NTL03PA05135 |
| UniParc | UPI00000C5F1C |
| UniProtKB Acc | Q9HU50 |
| UniProtKB ID | Q9HU50_PSEAE |
| UniRef100 | UniRef100_W0Z2V8 |
| UniRef50 | UniRef50_C5BJ27 |
| UniRef90 | UniRef90_A0A0A8RJL6 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
carboxyl-terminal processing protease, CtpA
|
| Synonyms | |
| Evidence for Translation |
Detected by MALDI-TOF/TOF (PMID:19333994).
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
Detected using LC-MS/MS (46.02 kDa, PMID:21751344)
|
| Charge (pH 7) | -4.29 |
| Kyte-Doolittle Hydrophobicity Value | -0.193 |
| Molecular Weight (kDa) | 46.0 |
| Isoelectric Point (pI) | 5.36 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7RPQ | HYDROLASE | 08/04/21 | Crystal Structure of carboxyl-terminal processing protease A, CtpA, of Pseudomonas aeruginosa | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 3.3 | X-RAY DIFFRACTION | 100.0 |
| 7RQH | HYDROLASE | 08/06/21 | Crystal Structure of carboxyl-terminal processing protease A mutant S302A, CtpA_S302A, of Pseudomonas aeruginosa | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 3.2 | X-RAY DIFFRACTION | 99.7 |
Virulence Evidence
| Source 1 | |
| Database | PseudoCAP |
| Category | |
| Evidence |
ECO:0000314
direct assay evidence used in manual assertion
|
| Host Organism | Cricetulus griseus |
| Notes | |
| Infection model | Cytotoxicity assay using chinese hamster ovary (CHO-K1) cells. |
| Reference | 24082078 |
| Source 2 | |
| Database | PseudoCAP |
| Category | |
| Evidence |
ECO:0000314
direct assay evidence used in manual assertion
|
| Host Organism | Mus musculus |
| Notes | |
| Infection model | Mouse model of acute pneumonia |
| Reference | 24082078 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 507 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG004726 (535 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA5134
Search term: ctpA
Search term: carboxyl-terminal processing protease, CtpA
|
Human Homologs
References
|
A carboxy-terminal processing protease gene is located immediately upstream of the invasion-associated locus from Bartonella bacilliformis.
Mitchell SJ, Minnick MF
Microbiology (Reading) 1997 Apr;143 ( Pt 4):1221-1233
PubMed ID: 9141685
|
|
Transport of CtpA protein from the cyanobacterium Synechocystis 6803 across the thylakoid membrane in chloroplasts.
Karnauchov I, Herrmann RG, Pakrasi HB, Klösgen RB
Eur J Biochem 1997 Oct 15;249(2):497-504
PubMed ID: 9370359
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|
|
Proteomic analysis of outer membrane vesicles derived from Pseudomonas aeruginosa.
Choi DS, Kim DK, Choi SJ, Lee J, Choi JP, Rho S, Park SH, Kim YK, Hwang D, Gho YS
Proteomics 2011 Aug;11(16):3424-9
PubMed ID: 21751344
|
|
The Pseudomonas aeruginosa periplasmic protease CtpA can affect systems that impact its ability to mount both acute and chronic infections.
Seo J, Darwin AJ
Infect Immun 2013 Dec;81(12):4561-70
PubMed ID: 24082078
|
|
Uncovering the structure and function of Pseudomonas aeruginosa periplasmic proteins by an in silico approach.
Caprari S, Brandi V, Pasquadibisceglie A, Polticelli F
J Biomol Struct Dyn 2020 Sep;38(15):4508-4520
PubMed ID: 31631799
|