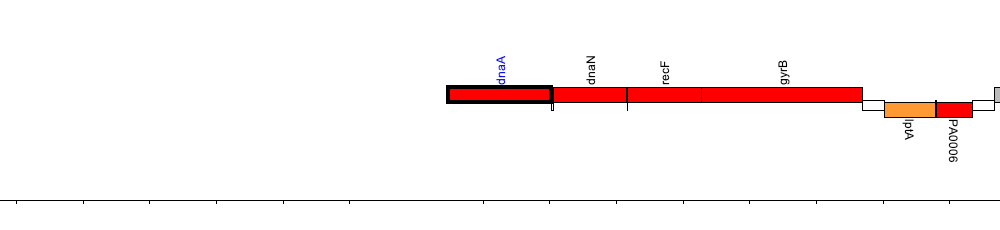
Pseudomonas aeruginosa PAO1, PA0001 (dnaA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0043565 | sequence-specific DNA binding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
10551881 | Reviewed by curator |
| Biological Process | GO:0006260 | DNA replication | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
12068813 | Reviewed by curator |
| Molecular Function | GO:0003677 | DNA binding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
7615570 | Reviewed by curator |
| Molecular Function | GO:0003688 | DNA replication origin binding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
19833870 | Reviewed by curator |
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006275 | regulation of DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00377
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006270 | DNA replication initiation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00377
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003688 | DNA replication origin binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00377
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00377
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00377
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR30050 | CHROMOSOMAL REPLICATION INITIATOR PROTEIN DNAA | - | - | 1 | 504 | 0.0 |
| SUPERFAMILY | SSF48295 | TrpR-like | IPR010921 | Trp repressor/replication initiator | 410 | 512 | 6.28E-37 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 178 | 344 | 1.2E-49 |
| FunFam | G3DSA:1.10.1750.10:FF:000001 | Chromosomal replication initiator protein DnaA | - | - | 421 | 514 | 1.4E-54 |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 211 | 420 | 2.3E-7 |
| Gene3D | G3DSA:3.30.300.180 | - | IPR038454 | DnaA, N-terminal domain superfamily | 2 | 102 | 1.3E-26 |
| CDD | cd00009 | AAA | - | - | 212 | 337 | 6.18022E-18 |
| Gene3D | G3DSA:1.10.8.60 | - | - | - | 345 | 415 | 6.2E-27 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 212 | 232 | 1.8E-52 |
| Pfam | PF08299 | Bacterial dnaA protein helix-turn-helix | IPR013159 | Chromosomal replication initiator, DnaA C-terminal | 423 | 491 | 1.8E-31 |
| FunFam | G3DSA:3.30.300.180:FF:000001 | Chromosomal replication initiator protein DnaA | - | - | 2 | 109 | 7.0E-41 |
| NCBIfam | TIGR00362 | JCVI: chromosomal replication initiator protein DnaA | IPR001957 | Chromosomal replication control, initiator DnaA | 6 | 512 | 0.0 |
| Pfam | PF00308 | Bacterial dnaA protein | IPR013317 | Chromosomal replication initiator protein DnaA | 179 | 396 | 1.0E-99 |
| FunFam | G3DSA:3.40.50.300:FF:000103 | Chromosomal replication initiator protein DnaA | - | - | 177 | 344 | 4.8E-100 |
| Pfam | PF11638 | DnaA N-terminal domain | IPR024633 | DnaA N-terminal domain | 4 | 63 | 2.7E-20 |
| CDD | cd06571 | Bac_DnaA_C | IPR013159 | Chromosomal replication initiator, DnaA C-terminal | 423 | 512 | 6.34824E-42 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 276 | 290 | 1.8E-52 |
| FunFam | G3DSA:1.10.8.60:FF:000003 | Chromosomal replication initiator protein DnaA | - | - | 345 | 415 | 2.4E-33 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 180 | 390 | 2.21E-44 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 472 | 491 | 1.8E-52 |
| SMART | SM00760 | bac_dnaa_c7seqb | IPR013159 | Chromosomal replication initiator, DnaA C-terminal | 422 | 491 | 9.1E-44 |
| Hamap | MF_00377 | Chromosomal replication initiator protein DnaA [dnaA]. | IPR001957 | Chromosomal replication control, initiator DnaA | 3 | 513 | 34.157234 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 310 | 337 | 1.8E-52 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 244 | 258 | 1.8E-52 |
| Gene3D | G3DSA:1.10.1750.10 | - | IPR010921 | Trp repressor/replication initiator | 421 | 514 | 4.0E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.