Pseudomonas aeruginosa PAO1, PA0156 (triA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
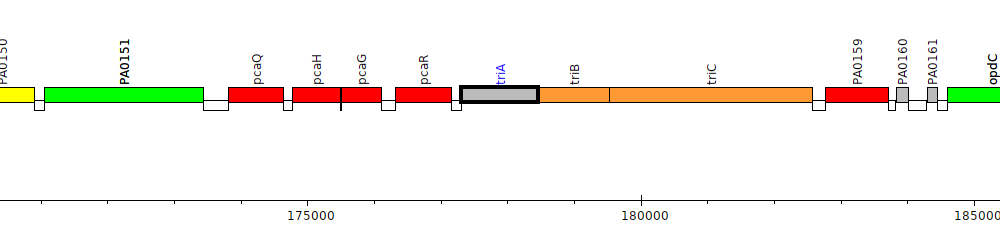
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA0156
|
| Name |
triA
Synonym: triA |
| Replicon | chromosome |
| Genomic location | 177307 - 178458 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_248846.1 |
| GI | 15595354 |
| Affymetrix | PA0156_at |
| Entrez | 882149 |
| GenBank | AAG03546.1 |
| INSDC | AAG03546.1 |
| NCBI Locus Tag | PA0156 |
| protein_id(GenBank) | gb|AAG03546.1|AE004453_9|gnl|PseudoCAP|PA0156 |
| TIGR | NTL03PA00157 |
| UniParc | UPI00000C4F6B |
| UniProtKB Acc | Q9I6X6 |
| UniProtKB ID | Q9I6X6_PSEAE |
| UniRef100 | UniRef100_U9HZG4 |
| UniRef50 | UniRef50_Q4KK51 |
| UniRef90 | UniRef90_W0YM83 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
Resistance-Nodulation-Cell Division (RND) triclosan efflux membrane fusion protein, TriA
|
| Synonyms |
probable RND efflux membrane fusion protein precursor Resistance-Nodulation-Cell Division (RND) triclosan efflux membrane fusion protein, TriA efflux membrane fusion protein precursor) |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -1.61 |
| Kyte-Doolittle Hydrophobicity Value | -0.137 |
| Molecular Weight (kDa) | 40.4 |
| Isoelectric Point (pI) | 6.53 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
AMR gene predictions from CARD database
Identified using the Resistance Gene Identifier (RGI)
| Model Name | Definition | Accession | Model Type (?) | SNP | Cutoff | Bit Score | Percent Identity | CARD Version |
| TriA | TriA is a membrane protein that is fused to TriB and both are required for the triclosan efflux pump function of TriABC-OpmH in P. aeruginosa. [PMID:17720796] | ARO:3003679 | protein homolog model | Perfect | 716.5 | 100.0 | 3.2.5 |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 6VEJ | TRANSPORT PROTEIN | 01/02/20 | TriABC transporter from Pseudomonas aeruginosa | Pseudomonas aeruginosa | 4.5 | ELECTRON MICROSCOPY | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 422 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000154 (998 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA0156
Search term: triA
|
Human Homologs
References
|
Identification and characterization of TriABC-OpmH, a triclosan efflux pump of Pseudomonas aeruginosa requiring two membrane fusion proteins.
Mima T, Joshi S, Gomez-Escalada M, Schweizer HP
J Bacteriol 2007 Nov;189(21):7600-9
PubMed ID: 17720796
|