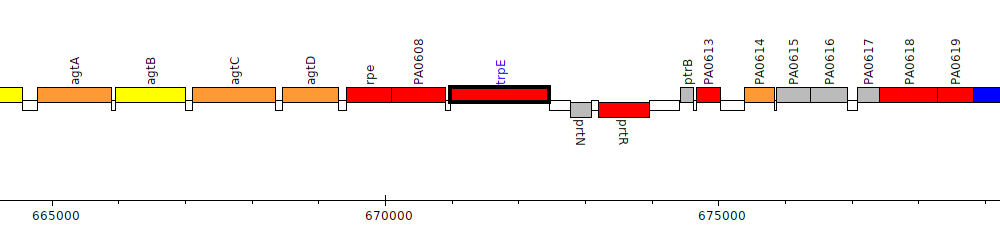
Pseudomonas aeruginosa PAO1, PA0609 (trpE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004049 | anthranilate synthase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
2153661 | Reviewed by curator |
| Biological Process | GO:0006558 | L-phenylalanine metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0000162 | tryptophan biosynthetic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
2153661 | Reviewed by curator |
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0006744 | ubiquinone biosynthetic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0000162 | tryptophan biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00564
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004049 | anthranilate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00564
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.60.120.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-6661 | 4-hydroxy-2(1H)-quinolone biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae02024 | Quorum sensing | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | ALL-CHORISMATE-PWY | superpathway of chorismate metabolism | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 4-hydroxy-2(1<i>H</i>)-quinolone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Ubiquinone biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | COMPLETE-ARO-PWY | superpathway of aromatic amino acid biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | TRPSYN-PWY | L-tryptophan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-6660 | 2-heptyl-3-hydroxy-4(1H)-quinolone biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| MetaCyc | acridone alkaloid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Phenylalanine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR11236 | AMINOBENZOATE/ANTHRANILATE SYNTHASE | IPR019999 | Anthranilate synthase component I-like | 13 | 490 | 0.0 |
| PRINTS | PR00095 | Anthranilate synthase component I signature | IPR019999 | Anthranilate synthase component I-like | 433 | 447 | 1.1E-25 |
| PRINTS | PR00095 | Anthranilate synthase component I signature | IPR019999 | Anthranilate synthase component I-like | 418 | 432 | 1.1E-25 |
| PRINTS | PR00095 | Anthranilate synthase component I signature | IPR019999 | Anthranilate synthase component I-like | 338 | 351 | 1.1E-25 |
| SUPERFAMILY | SSF56322 | ADC synthase | IPR005801 | ADC synthase | 14 | 485 | 0.0 |
| PRINTS | PR00095 | Anthranilate synthase component I signature | IPR019999 | Anthranilate synthase component I-like | 324 | 337 | 1.1E-25 |
| Pfam | PF00425 | chorismate binding enzyme | IPR015890 | Chorismate-utilising enzyme, C-terminal | 223 | 476 | 4.9E-91 |
| NCBIfam | TIGR00564 | JCVI: anthranilate synthase component I | IPR005256 | Anthranilate synthase component I, PabB-like | 26 | 484 | 0.0 |
| Gene3D | G3DSA:3.60.120.10 | Anthranilate synthase | IPR005801 | ADC synthase | 1 | 491 | 0.0 |
| Pfam | PF04715 | Anthranilate synthase component I, N terminal region | IPR006805 | Anthranilate synthase component I, N-terminal | 27 | 168 | 8.3E-31 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.