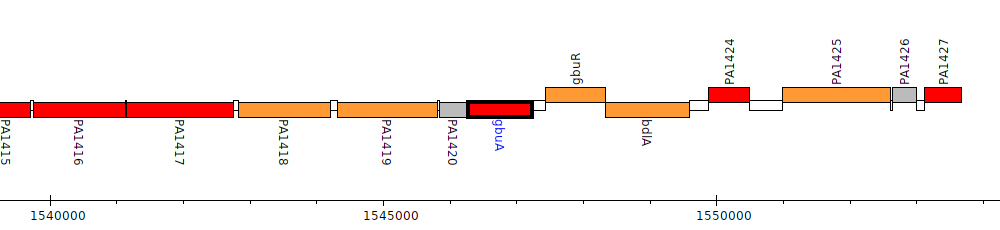
Pseudomonas aeruginosa PAO1, PA1421 (gbuA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006527 | arginine catabolic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
3141581 | Reviewed by curator |
| Molecular Function | GO:0047971 | guanidinobutyrase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
3141581 | Reviewed by curator |
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11358
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | PWY-40 | putrescine biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00330 | Arginine and proline metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY0-823 | L-arginine degradation III (arginine decarboxylase/agmatinase pathway) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | ARGDEG-PWY | superpathway of L-arginine, putrescine, and 4-aminobutyrate degradation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Arginine and proline metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | POLYAMSYN-PWY | superpathway of polyamine biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | PWY-5742 | L-arginine degradation IX (arginine:pyruvate transaminase pathway) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.800.10 | Ureohydrolase domain | - | - | 1 | 319 | 1.4E-114 |
| PRINTS | PR00116 | Arginase signature | IPR006035 | Ureohydrolase | 238 | 267 | 2.2E-13 |
| PANTHER | PTHR11358 | ARGINASE/AGMATINASE | IPR006035 | Ureohydrolase | 13 | 314 | 3.0E-102 |
| NCBIfam | TIGR01230 | JCVI: agmatinase | IPR005925 | Agmatinase-related | 34 | 311 | 1.2E-62 |
| SUPERFAMILY | SSF52768 | Arginase/deacetylase | IPR023696 | Ureohydrolase domain superfamily | 4 | 313 | 5.94E-100 |
| PRINTS | PR00116 | Arginase signature | IPR006035 | Ureohydrolase | 125 | 140 | 2.2E-13 |
| PIRSF | PIRSF036979 | Arginase | IPR006035 | Ureohydrolase | 15 | 316 | 2.2E-102 |
| Pfam | PF00491 | Arginase family | IPR006035 | Ureohydrolase | 39 | 310 | 1.1E-87 |
| CDD | cd11592 | Agmatinase_PAH | - | - | 19 | 310 | 0.0 |
| FunFam | G3DSA:3.40.800.10:FF:000002 | Agmatinase | - | - | 2 | 319 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.