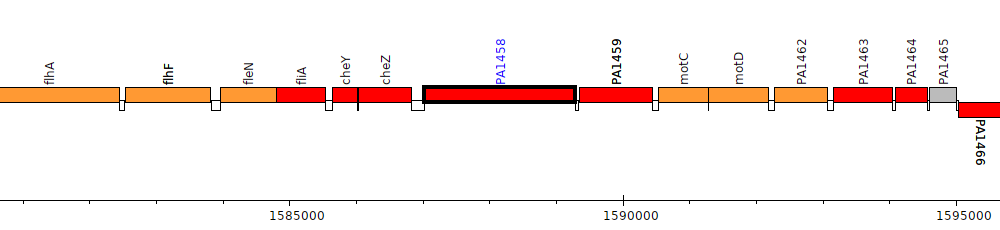
Pseudomonas aeruginosa PAO1, PA1458
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.20.120.160
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004673 | protein histidine kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM01231
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:SM01231
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.287.560
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.287.560
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Two-component System |
ECO:0000037
not_recorded |
|||
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Chemotaxis |
ECO:0000037
not_recorded |
|||
| KEGG | pae02030 | Bacterial chemotaxis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | Coil | - | - | 14 | 34 | - |
| SUPERFAMILY | SSF47226 | Histidine-containing phosphotransfer domain, HPT domain | IPR036641 | HPT domain superfamily | 6 | 128 | 3.14E-33 |
| SMART | SM00073 | hpt_2 | IPR008207 | Signal transduction histidine kinase, phosphotransfer (Hpt) domain | 6 | 103 | 3.0E-19 |
| PANTHER | PTHR43395 | SENSOR HISTIDINE KINASE CHEA | - | - | 6 | 751 | 0.0 |
| SMART | SM01231 | H_kinase_dim_2 | IPR004105 | Histidine kinase CheA-like, homodimeric domain | 367 | 427 | 9.7E-19 |
| FunFam | G3DSA:3.30.565.10:FF:000016 | Chemotaxis protein CheA, putative | - | - | 428 | 614 | 5.7E-87 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 129 | 164 | - |
| SMART | SM00260 | chew_2 | IPR002545 | CheW-like domain | 607 | 746 | 8.6E-43 |
| SUPERFAMILY | SSF55874 | ATPase domain of HSP90 chaperone/DNA topoisomerase II/histidine kinase | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 430 | 613 | 7.47E-39 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 599 | 612 | 4.1E-12 |
| FunFam | G3DSA:1.20.120.160:FF:000008 | Chemotaxis sensor histidine kinase CheA | - | - | 6 | 138 | 3.0E-61 |
| Pfam | PF01584 | CheW-like domain | IPR002545 | CheW-like domain | 621 | 748 | 6.4E-24 |
| Gene3D | G3DSA:1.20.120.160 | HPT domain | IPR036641 | HPT domain superfamily | 4 | 138 | 2.4E-36 |
| Gene3D | G3DSA:2.30.30.40 | SH3 Domains | - | - | 685 | 746 | 7.7E-22 |
| FunFam | G3DSA:2.30.30.40:FF:000048 | Chemotaxis protein CheA, putative | - | - | 684 | 746 | 2.7E-29 |
| CDD | cd00088 | HPT | IPR008207 | Signal transduction histidine kinase, phosphotransfer (Hpt) domain | 6 | 104 | 2.37121E-19 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 517 | 531 | 4.1E-12 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 135 | 154 | - |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 575 | 593 | 4.1E-12 |
| Pfam | PF02895 | Signal transducing histidine kinase, homodimeric domain | IPR004105 | Histidine kinase CheA-like, homodimeric domain | 368 | 427 | 6.3E-14 |
| Gene3D | G3DSA:1.10.287.560 | - | IPR037006 | Histidine kinase CheA-like, homodimeric domain superfamily | 368 | 427 | 3.3E-17 |
| SMART | SM00387 | HKATPase_4 | IPR003594 | Histidine kinase/HSP90-like ATPase | 472 | 615 | 1.0E-24 |
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 475 | 614 | 1.4E-16 |
| CDD | cd16916 | HATPase_CheA-like | - | - | 435 | 612 | 5.17471E-93 |
| SUPERFAMILY | SSF47384 | Homodimeric domain of signal transducing histidine kinase | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 368 | 428 | 5.89E-14 |
| SUPERFAMILY | SSF50341 | CheW-like | IPR036061 | CheW-like domain superfamily | 614 | 746 | 5.36E-32 |
| CDD | cd00731 | CheA_reg | - | - | 615 | 745 | 2.58715E-48 |
| Gene3D | G3DSA:3.30.565.10 | - | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 428 | 614 | 2.2E-54 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 559 | 569 | 4.1E-12 |
| Pfam | PF01627 | Hpt domain | IPR008207 | Signal transduction histidine kinase, phosphotransfer (Hpt) domain | 8 | 100 | 8.9E-18 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 264 | 298 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.