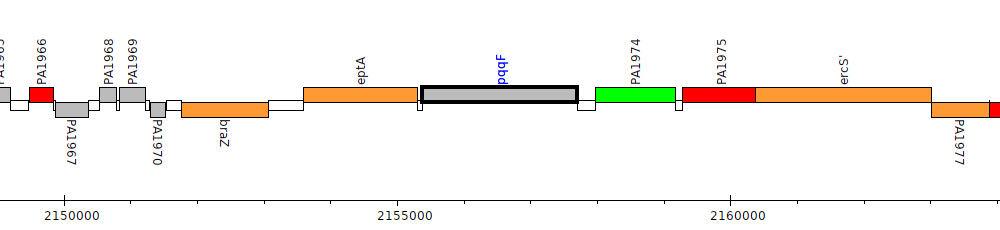
Pseudomonas aeruginosa PAO1, PA1973 (pqqF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0051188 | obsolete cofactor biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008237 | metallopeptidase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: RefSeq:YP_002963390.1
|
ECO:0000250 sequence similarity evidence used in manual assertion |
8606199 | Reviewed by curator |
| Cellular Component | GO:0042597 | periplasmic space |
Inferred from Sequence or Structural Similarity
Term mapped from: RefSeq:YP_002963390.1
|
ECO:0000250 sequence similarity evidence used in manual assertion |
8606199 | Reviewed by curator |
| Biological Process | GO:0018189 | pyrroloquinoline quinone biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
7665488 | Reviewed by curator |
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0018189 | pyrroloquinoline quinone biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF63411
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Pyrroloquinoline quinone biosynthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF63411 | LuxS/MPP-like metallohydrolase | IPR011249 | Metalloenzyme, LuxS/M16 peptidase-like | 12 | 218 | 4.68E-35 |
| Gene3D | G3DSA:3.30.830.10 | - | - | - | 2 | 228 | 6.1E-50 |
| Pfam | PF05193 | Peptidase M16 inactive domain | IPR007863 | Peptidase M16, C-terminal | 184 | 341 | 7.0E-12 |
| Gene3D | G3DSA:3.30.830.10 | - | - | - | 593 | 770 | 2.5E-20 |
| SUPERFAMILY | SSF63411 | LuxS/MPP-like metallohydrolase | IPR011249 | Metalloenzyme, LuxS/M16 peptidase-like | 591 | 768 | 1.15E-18 |
| Pfam | PF00675 | Insulinase (Peptidase family M16) | IPR011765 | Peptidase M16, N-terminal | 24 | 144 | 1.5E-25 |
| NCBIfam | TIGR02110 | JCVI: pyrroloquinoline quinone biosynthesis protein PqqF | IPR011844 | Coenzyme PQQ biosynthesis protein PqqF | 12 | 688 | 0.0 |
| PANTHER | PTHR43690 | NARDILYSIN | - | - | 633 | 730 | 2.3E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.