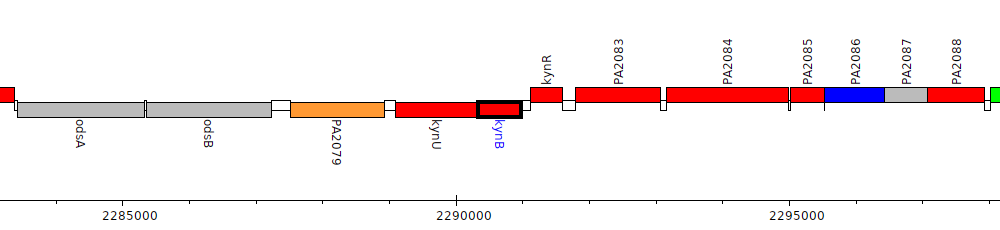
Pseudomonas aeruginosa PAO1, PA2081 (kynB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008150 | biological_process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004061 | arylformamidase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
14592712 | Reviewed by curator |
| Biological Process | GO:0019441 | tryptophan catabolic process to kynurenine | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
17337571 | Reviewed by curator |
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Cellular Component | GO:0005575 | cellular_component | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004328 | formamidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01969
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019441 | tryptophan catabolic process to kynurenine |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.50.30.50
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004061 | arylformamidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.50.30.50
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-5651 | L-tryptophan degradation to 2-amino-3-carboxymuconate semialdehyde | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | anthranilate |
ECO:0000037
not_recorded |
|||
| PseudoCyc | TRYPTOPHAN-DEGRADATION-1 | L-tryptophan degradation III (eukaryotic) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00380 | Tryptophan metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | kynurenine pathway |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.50.30.50 | Putative cyclase | IPR037175 | Kynurenine formamidase superfamily | 1 | 213 | 1.4E-67 |
| Hamap | MF_01969 | Kynurenine formamidase [kynB]. | IPR017484 | Kynurenine formamidase, bacteria | 5 | 209 | 45.107018 |
| FunFam | G3DSA:3.50.30.50:FF:000001 | Kynurenine formamidase | - | - | 3 | 209 | 5.0E-89 |
| PANTHER | PTHR31118 | CYCLASE-LIKE PROTEIN 2 | IPR007325 | Kynurenine formamidase/cyclase-like | 5 | 206 | 9.5E-25 |
| SUPERFAMILY | SSF102198 | Putative cyclase | IPR037175 | Kynurenine formamidase superfamily | 4 | 207 | 9.81E-60 |
| NCBIfam | TIGR03035 | JCVI: arylformamidase | IPR017484 | Kynurenine formamidase, bacteria | 5 | 209 | 3.7E-104 |
| Pfam | PF04199 | Putative cyclase | IPR007325 | Kynurenine formamidase/cyclase-like | 8 | 152 | 6.9E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.