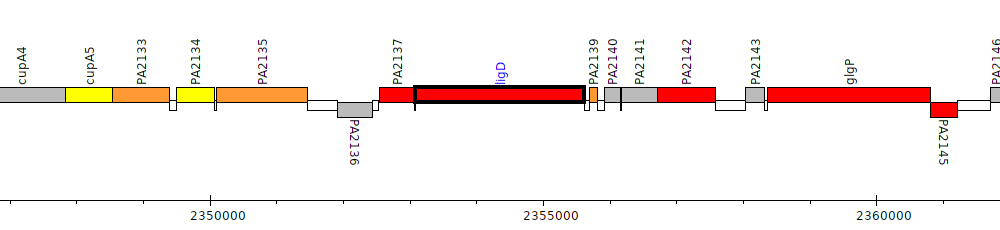
Pseudomonas aeruginosa PAO1, PA2138 (ligD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016779 | nucleotidyltransferase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15520014 | Reviewed by curator |
| Molecular Function | GO:0003910 | DNA ligase (ATP) activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15520014 | Reviewed by curator |
| Molecular Function | GO:0003887 | DNA-directed DNA polymerase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15520014 | Reviewed by curator |
| Molecular Function | GO:0004532 | exoribonuclease activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15897197 | Reviewed by curator |
| Biological Process | GO:0090503 | RNA phosphodiester bond hydrolysis, exonucleolytic | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15897197 | Reviewed by curator |
| Biological Process | GO:0071897 | DNA biosynthetic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15520014 | Reviewed by curator |
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:PF01068
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003910 | DNA ligase (ATP) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01068
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006310 | DNA recombination |
Inferred from Sequence Model
Term mapped from: InterPro:PF01068
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01068
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae03450 | Non-homologous end-joining | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR02776 | JCVI: DNA ligase D | IPR014143 | DNA ligase D | 228 | 823 | 0.0 |
| Gene3D | G3DSA:3.30.1490.70 | - | - | - | 229 | 397 | 1.2E-46 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 514 | 546 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 189 | 211 | - |
| CDD | cd04862 | PaeLigD_Pol_like | IPR033651 | LigD polymerase domain, PaeLigD-type | 566 | 792 | 1.18596E-126 |
| Gene3D | G3DSA:3.30.470.30 | DNA ligase/mRNA capping enzyme | - | - | 241 | 356 | 1.2E-46 |
| Pfam | PF01068 | ATP dependent DNA ligase domain | IPR012310 | DNA ligase, ATP-dependent, central | 224 | 399 | 1.9E-22 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 8 | 32 | - |
| PANTHER | PTHR42705 | BIFUNCTIONAL NON-HOMOLOGOUS END JOINING PROTEIN LIGD | - | - | 228 | 836 | 3.6E-111 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 403 | 523 | 3.98E-28 |
| CDD | cd07906 | Adenylation_DNA_ligase_LigD_LigC | - | - | 217 | 400 | 8.28624E-80 |
| NCBIfam | TIGR02779 | JCVI: non-homologous end-joining DNA ligase, ligase domain | IPR014146 | DNA ligase D, ligase domain | 219 | 521 | 2.2E-110 |
| NCBIfam | TIGR02777 | JCVI: DNA ligase D, 3'-phosphoesterase domain | IPR014144 | DNA ligase D, 3'-phosphoesterase domain | 7 | 162 | 1.5E-70 |
| CDD | cd07971 | OBF_DNA_ligase_LigD | - | - | 403 | 520 | 1.96759E-49 |
| NCBIfam | TIGR02778 | JCVI: non-homologous end-joining DNA ligase, polymerase domain | IPR014145 | DNA ligase D, polymerase domain | 549 | 793 | 2.7E-88 |
| Gene3D | G3DSA:3.90.920.10 | DNA primase, PRIM domain | - | - | 556 | 840 | 2.7E-92 |
| SUPERFAMILY | SSF56091 | DNA ligase/mRNA capping enzyme, catalytic domain | - | - | 216 | 399 | 2.3E-45 |
| Pfam | PF04679 | ATP dependent DNA ligase C terminal region | IPR012309 | DNA ligase, ATP-dependent, C-terminal | 418 | 515 | 1.2E-23 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 32 | - |
| Gene3D | G3DSA:2.40.50.140 | - | IPR012340 | Nucleic acid-binding, OB-fold | 402 | 523 | 8.1E-37 |
| Pfam | PF13298 | DNA polymerase Ligase (LigD) | IPR014144 | DNA ligase D, 3'-phosphoesterase domain | 40 | 144 | 5.1E-39 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.