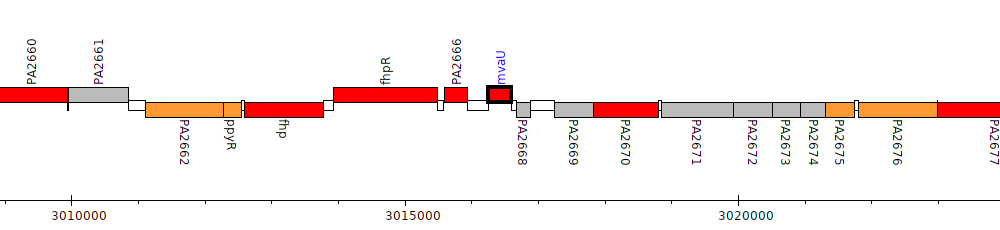
Pseudomonas aeruginosa PAO1, PA2667 (mvaU)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0000976 | transcription regulatory region sequence-specific DNA binding | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
19684136 | Reviewed by curator |
| Biological Process | GO:1900377 | negative regulation of secondary metabolite biosynthetic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
19684136 | Reviewed by curator |
| Biological Process | GO:0010963 | regulation of L-arginine import | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
19684136 | Reviewed by curator |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF00816
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030527 | structural constituent of chromatin |
Inferred from Sequence Model
Term mapped from: InterPro:PF00816
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00816
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd16170 | MvaT_DBD | IPR035616 | MvaT, DNA-binding domain | 75 | 116 | 2.05925E-21 |
| Coils | Coil | Coil | - | - | 4 | 27 | - |
| Pfam | PF00816 | H-NS histone family | IPR001801 | Histone-like protein H-NS | 13 | 105 | 1.8E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.