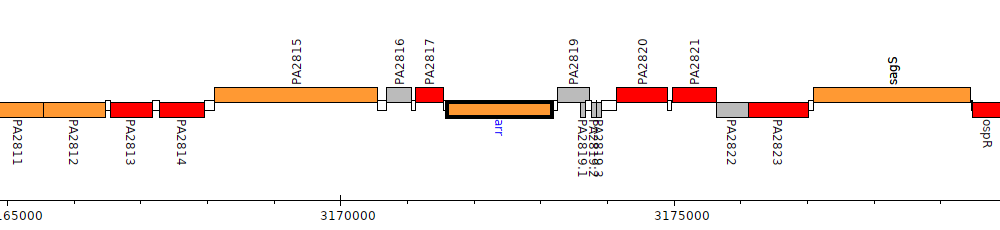
Pseudomonas aeruginosa PAO1, PA2818 (arr)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:1900190 | regulation of single-species biofilm formation | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
16121184 | Reviewed by curator |
| Molecular Function | GO:0004114 | 3',5'-cyclic-nucleotide phosphodiesterase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
16121184 | Reviewed by curator |
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
16121184 | Reviewed by curator |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR33121 | CYCLIC DI-GMP PHOSPHODIESTERASE PDEF | - | - | 220 | 510 | 1.2E-85 |
| CDD | cd01948 | EAL | IPR001633 | EAL domain | 267 | 505 | 2.51117E-85 |
| SUPERFAMILY | SSF141868 | EAL domain-like | IPR035919 | EAL domain superfamily | 267 | 513 | 8.11E-76 |
| Gene3D | G3DSA:3.20.20.450 | EAL domain | IPR035919 | EAL domain superfamily | 261 | 522 | 4.1E-81 |
| Pfam | PF00563 | EAL domain | IPR001633 | EAL domain | 270 | 500 | 6.9E-65 |
| SMART | SM00052 | duf2_2 | IPR001633 | EAL domain | 261 | 505 | 6.1E-89 |
| Pfam | PF12792 | CSS motif domain associated with EAL | IPR024744 | Putative cyclic diguanylate phosphodiesterase, CSS motif-containing domain | 46 | 234 | 3.3E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.