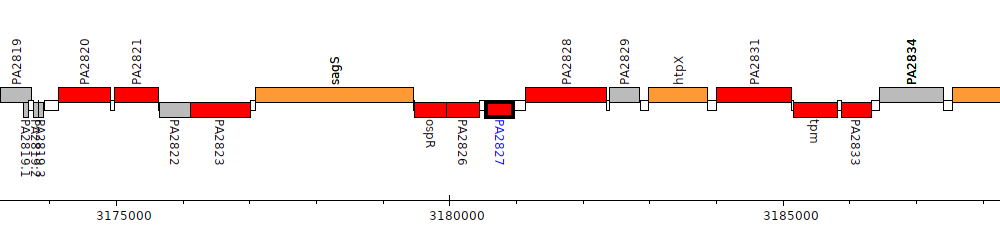
Pseudomonas aeruginosa PAO1, PA2827
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:1901530 | response to hypochlorite | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
23687271 | Reviewed by curator |
| Biological Process | GO:0034599 | cellular response to oxidative stress | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
23687271 | Reviewed by curator |
| Molecular Function | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
23687271 | Reviewed by curator |
| Biological Process | GO:0009405 | pathogenesis | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
23687271 | Reviewed by curator |
| Molecular Function | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR10173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR10173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01641
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0030091 | protein repair |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR10173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:2.170.150.20:FF:000001 | Peptide methionine sulfoxide reductase MsrB | - | - | 1 | 132 | 6.7E-64 |
| SUPERFAMILY | SSF51316 | Mss4-like | IPR011057 | Mss4-like superfamily | 7 | 131 | 1.08E-56 |
| Pfam | PF01641 | SelR domain | IPR002579 | Peptide methionine sulphoxide reductase MrsB domain | 11 | 128 | 1.4E-52 |
| Gene3D | G3DSA:2.170.150.20 | Peptide methionine sulfoxide reductase. | - | - | 2 | 132 | 1.3E-59 |
| PANTHER | PTHR10173 | METHIONINE SULFOXIDE REDUCTASE | IPR028427 | Peptide methionine sulfoxide reductase MsrB | 9 | 131 | 6.1E-61 |
| NCBIfam | TIGR00357 | JCVI: peptide-methionine (R)-S-oxide reductase MsrB | IPR002579 | Peptide methionine sulphoxide reductase MrsB domain | 9 | 131 | 1.4E-53 |
| Hamap | MF_01400 | Peptide methionine sulfoxide reductase MsrB [msrB]. | IPR002579 | Peptide methionine sulphoxide reductase MrsB domain | 7 | 130 | 37.608044 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.