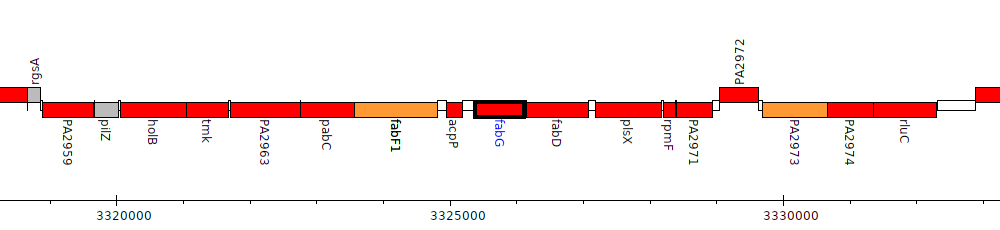
Pseudomonas aeruginosa PAO1, PA2967 (fabG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044255 | cellular lipid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004316 | 3-oxoacyl-[acyl-carrier-protein] reductase (NADPH) activity | Inferred from Genetic Interaction | ECO:0000012 functional complementation evidence |
10781572 | Reviewed by curator |
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01830
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004316 | 3-oxoacyl-[acyl-carrier-protein] reductase (NADPH) activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01830
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006633 | fatty acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01830
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00780 | Biotin metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00061 | Fatty acid biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01212 | Fatty acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01040 | Biosynthesis of unsaturated fatty acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.40.50.720:FF:000037 | 3-oxoacyl-[acyl-carrier-protein] reductase FabG | - | - | 1 | 247 | 1.1E-127 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 128 | 144 | 3.6E-46 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 154 | 173 | 3.3E-12 |
| NCBIfam | TIGR01830 | JCVI: 3-oxoacyl-[acyl-carrier-protein] reductase | IPR011284 | 3-oxoacyl-(acyl-carrier-protein) reductase | 8 | 245 | 3.9E-100 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 154 | 173 | 3.6E-46 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 81 | 92 | 3.3E-12 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 81 | 92 | 3.6E-46 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 1 | 247 | 4.8E-87 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 7 | 24 | 3.6E-46 |
| PANTHER | PTHR42879 | 3-OXOACYL-(ACYL-CARRIER-PROTEIN) REDUCTASE | - | - | 1 | 245 | 5.1E-85 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 208 | 228 | 3.6E-46 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 175 | 192 | 3.6E-46 |
| SMART | SM00822 | This enzymatic domain is part of bacterial polyketide synthases and catalyses the first step in the reductive modification of the beta-carbonyl centres in the growing polyketide chain. It uses NADPH to reduce the keto group to a hydroxy group. | - | - | 6 | 185 | 1.3E-5 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 134 | 142 | 3.3E-12 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 4 | 246 | 6.59E-83 |
| CDD | cd05333 | BKR_SDR_c | - | - | 6 | 245 | 7.55475E-125 |
| Pfam | PF13561 | Enoyl-(Acyl carrier protein) reductase | - | - | 12 | 244 | 2.9E-63 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.