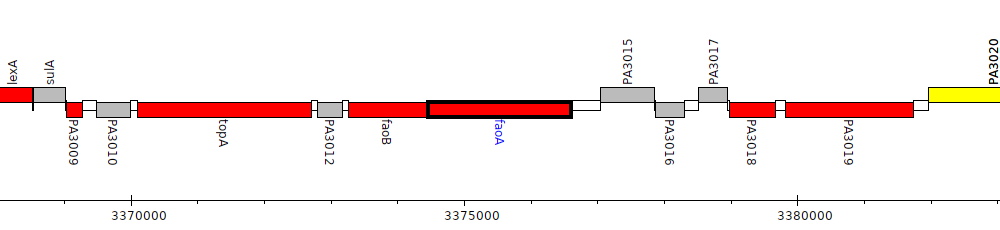
Pseudomonas aeruginosa PAO1, PA3014 (faoA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050695 | benzoylformate decarboxylase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0019605 | butyrate metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006568 | tryptophan metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006574 | valine catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0019541 | propionate metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006554 | lysine catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0019482 | beta-alanine metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006631 | fatty acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006633 | fatty acid biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006552 | leucine catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006550 | isoleucine catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044255 | cellular lipid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008692 | 3-hydroxybutyryl-CoA epimerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0070403 | NAD+ binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02737
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006631 | fatty acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0036125 | fatty acid beta-oxidation multienzyme complex |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009062 | fatty acid catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004165 | dodecenoyl-CoA delta-isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004300 | enoyl-CoA hydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Fatty acid metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Lysine degradation |
ECO:0000037
not_recorded |
|||
| KEGG | pae00903 | Limonene and pinene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00930 | Caprolactam degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00410 | beta-Alanine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00281 | Geraniol degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00650 | Butanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01212 | Fatty acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Propanoate metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00362 | Benzoate degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Valine, leucine and isoleucine degradation |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Beta-Alanine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00280 | Valine, leucine and isoleucine degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Fatty acid biosynthesis (path 2) |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Tryptophan metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Butanoate metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00640 | Propanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00380 | Tryptophan metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01040 | Biosynthesis of unsaturated fatty acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00310 | Lysine degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00071 | Fatty acid degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00725 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal domain | IPR006108 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal | 497 | 592 | 3.3E-25 |
| CDD | cd06558 | crotonase-like | - | - | 16 | 204 | 6.04648E-57 |
| NCBIfam | TIGR02437 | JCVI: fatty acid oxidation complex subunit alpha FadB | IPR012799 | Fatty oxidation complex, alpha subunit FadB | 1 | 714 | 0.0 |
| Pfam | PF00378 | Enoyl-CoA hydratase/isomerase | IPR001753 | Enoyl-CoA hydratase/isomerase | 17 | 212 | 9.0E-40 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 314 | 496 | 1.76E-56 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 314 | 499 | 2.4E-66 |
| PANTHER | PTHR43612 | TRIFUNCTIONAL ENZYME SUBUNIT ALPHA | - | - | 3 | 714 | 0.0 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 496 | 616 | 3.57E-29 |
| Gene3D | G3DSA:3.90.226.10 | - | - | - | 1 | 313 | 8.0E-121 |
| SUPERFAMILY | SSF52096 | ClpP/crotonase | IPR029045 | ClpP/crotonase-like domain superfamily | 2 | 308 | 8.83E-70 |
| Gene3D | G3DSA:1.10.1040.50 | - | - | - | 500 | 704 | 6.8E-71 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 622 | 714 | 6.29E-24 |
| Hamap | MF_01621 | Fatty acid oxidation complex subunit alpha [fadB]. | IPR012799 | Fatty oxidation complex, alpha subunit FadB | 1 | 715 | 52.228764 |
| FunFam | G3DSA:3.40.50.720:FF:000009 | Fatty oxidation complex, alpha subunit | - | - | 314 | 499 | 5.6E-73 |
| FunFam | G3DSA:1.10.1040.50:FF:000001 | Fatty acid oxidation complex subunit alpha | - | - | 500 | 714 | 2.3E-118 |
| FunFam | G3DSA:3.90.226.10:FF:000018 | Fatty acid oxidation complex subunit alpha | - | - | 1 | 313 | 0.0 |
| Pfam | PF02737 | 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain | IPR006176 | 3-hydroxyacyl-CoA dehydrogenase, NAD binding | 317 | 495 | 8.5E-61 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.