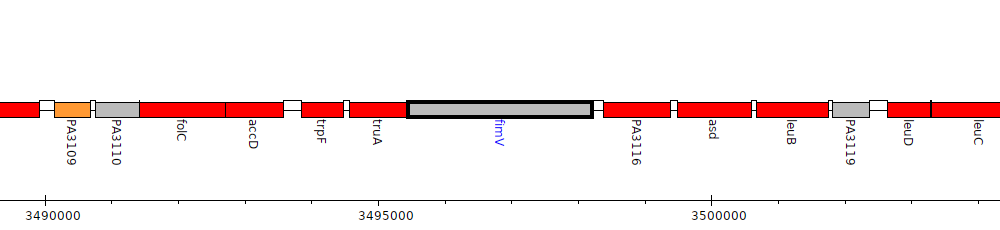
Pseudomonas aeruginosa PAO1, PA3115 (fimV)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0042834 | peptidoglycan binding | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
21097635 | Reviewed by curator |
| Cellular Component | GO:0044096 | type IV pilus | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
21097635 | Reviewed by curator |
| Cellular Component | GO:0015627 | type II protein secretion system complex | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
21527471 | Reviewed by curator |
| Biological Process | GO:0006928 | movement of cell or subcellular component | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.25.40.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.10.350.10 | LysM domain | IPR036779 | LysM domain superfamily | 163 | 238 | 1.6E-10 |
| Gene3D | G3DSA:1.25.40.10 | Tetratricopeptide repeat domain | IPR011990 | Tetratricopeptide-like helical domain superfamily | 547 | 626 | 3.9E-6 |
| PANTHER | PTHR18898 | NUCLEOPROTEIN TPR-RELATED | - | - | 327 | 898 | 8.4E-10 |
| CDD | cd00118 | LysM | IPR018392 | LysM domain | 175 | 229 | 3.23679E-6 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 237 | 312 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 269 | 283 | - |
| SMART | SM00257 | LysM_2 | IPR018392 | LysM domain | 175 | 230 | 0.0037 |
| SUPERFAMILY | SSF48452 | TPR-like | IPR011990 | Tetratricopeptide-like helical domain superfamily | 551 | 607 | 2.85E-5 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 380 | 394 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 785 | 816 | - |
| Gene3D | G3DSA:1.20.58.2200 | - | IPR038440 | FimV, C-terminal domain superfamily | 858 | 919 | 1.9E-27 |
| NCBIfam | TIGR03505 | JCVI: FimV/HubP N-terminal domain | IPR020012 | Motility protein FimV, N-terminal | 182 | 254 | 4.0E-30 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 237 | 259 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 411 | 445 | - |
| Coils | Coil | Coil | - | - | 319 | 367 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 372 | 445 | - |
| Coils | Coil | Coil | - | - | 904 | 919 | - |
| NCBIfam | TIGR03504 | JCVI: FimV/HubP C-terminal domain | IPR020011 | Motility protein FimV, C-terminal | 875 | 918 | 6.6E-21 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 140 | 177 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.