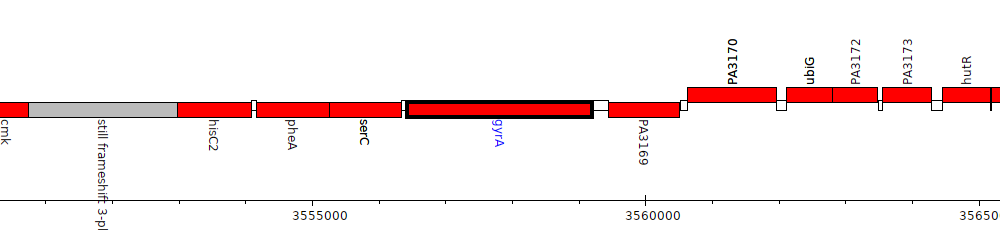
Pseudomonas aeruginosa PAO1, PA3168 (gyrA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006259 | DNA metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004347 | glucose-6-phosphate isomerase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Cellular Component | GO:0005694 | chromosome |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01897
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006259 | DNA metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.90.199.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003918 | DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006265 | DNA topological change |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003916 | DNA topoisomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF03989
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | Coil | - | - | 500 | 520 | - |
| SUPERFAMILY | SSF56719 | Type II DNA topoisomerase | IPR013760 | DNA topoisomerase, type IIA-like domain superfamily | 30 | 523 | 0.0 |
| PANTHER | PTHR43493 | DNA GYRASE/TOPOISOMERASE SUBUNIT A | - | - | 4 | 887 | 0.0 |
| Pfam | PF00521 | DNA gyrase/topoisomerase IV, subunit A | IPR002205 | DNA topoisomerase, type IIA, domain A | 32 | 507 | 0.0 |
| SUPERFAMILY | SSF101904 | GyrA/ParC C-terminal domain-like | IPR035516 | DNA gyrase/topoisomerase IV, subunit A, C-terminal | 534 | 898 | 1.7E-99 |
| Gene3D | G3DSA:3.30.1360.40 | - | - | - | 236 | 334 | 0.0 |
| FunFam | G3DSA:3.30.1360.40:FF:000002 | DNA gyrase subunit A | - | - | 236 | 334 | 2.0E-41 |
| Gene3D | G3DSA:3.90.199.10 | Topoisomerase II, domain 5 | IPR013758 | DNA topoisomerase, type IIA, domain A, alpha-beta | 32 | 522 | 0.0 |
| Pfam | PF03989 | DNA gyrase C-terminal domain, beta-propeller | IPR006691 | DNA gyrase/topoisomerase IV, subunit A, C-terminal repeat | 737 | 781 | 1.9E-13 |
| Pfam | PF03989 | DNA gyrase C-terminal domain, beta-propeller | IPR006691 | DNA gyrase/topoisomerase IV, subunit A, C-terminal repeat | 837 | 883 | 2.4E-12 |
| Gene3D | G3DSA:2.120.10.90 | - | IPR035516 | DNA gyrase/topoisomerase IV, subunit A, C-terminal | 532 | 885 | 9.2E-127 |
| SMART | SM00434 | topIV4 | IPR002205 | DNA topoisomerase, type IIA, domain A | 11 | 500 | 0.0 |
| Pfam | PF03989 | DNA gyrase C-terminal domain, beta-propeller | IPR006691 | DNA gyrase/topoisomerase IV, subunit A, C-terminal repeat | 539 | 586 | 1.5E-8 |
| Pfam | PF03989 | DNA gyrase C-terminal domain, beta-propeller | IPR006691 | DNA gyrase/topoisomerase IV, subunit A, C-terminal repeat | 591 | 638 | 1.6E-8 |
| Pfam | PF03989 | DNA gyrase C-terminal domain, beta-propeller | IPR006691 | DNA gyrase/topoisomerase IV, subunit A, C-terminal repeat | 689 | 732 | 2.8E-8 |
| Pfam | PF03989 | DNA gyrase C-terminal domain, beta-propeller | IPR006691 | DNA gyrase/topoisomerase IV, subunit A, C-terminal repeat | 788 | 833 | 2.2E-12 |
| NCBIfam | TIGR01063 | JCVI: DNA gyrase subunit A | - | - | 8 | 638 | 0.0 |
| FunFam | G3DSA:3.90.199.10:FF:000001 | DNA gyrase subunit A | - | - | 32 | 251 | 8.0E-113 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 890 | 923 | - |
| Hamap | MF_01897 | DNA gyrase subunit A [gyrA]. | IPR005743 | DNA gyrase, subunit A | 1 | 885 | 23.955879 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 881 | 923 | - |
| CDD | cd00187 | TOP4c | IPR002205 | DNA topoisomerase, type IIA, domain A | 30 | 510 | 0.0 |
| Gene3D | G3DSA:1.10.268.10 | Topoisomerase, domain 3 | IPR013757 | DNA topoisomerase, type IIA, alpha-helical domain superfamily | 371 | 493 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.