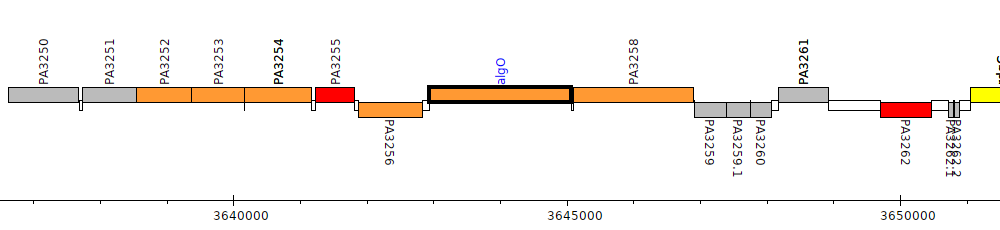
Pseudomonas aeruginosa PAO1, PA3257 (algO)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0042493 | response to drug | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
1447154 | Reviewed by curator |
| Biological Process | GO:0007165 | signal transduction | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
24373018 | Reviewed by curator |
| Molecular Function | GO:0004252 | serine-type endopeptidase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PP_1719
|
ECO:0000250 sequence similarity evidence used in manual assertion |
24373018 | Reviewed by curator |
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008236 | serine-type peptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00245
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00595
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:SM00245
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 657 | 687 | - |
| Gene3D | G3DSA:3.90.226.10 | - | - | - | 362 | 553 | 8.6E-80 |
| FunFam | G3DSA:3.90.226.10:FF:000090 | Tail-specific protease | - | - | 346 | 575 | 1.4E-77 |
| Pfam | PF17804 | Tail specific protease N-terminal domain | IPR040573 | Tail specific protease, N-terminal domain | 63 | 250 | 1.9E-65 |
| Pfam | PF03572 | Peptidase family S41 | IPR005151 | Tail specific protease | 381 | 554 | 1.9E-51 |
| Pfam | PF11818 | C-terminal domain of tail specific protease (DUF3340) | IPR020992 | Tail specific protease, C-terminal | 560 | 700 | 6.9E-37 |
| SMART | SM00245 | tsp_4 | IPR005151 | Tail specific protease | 347 | 556 | 3.1E-56 |
| SUPERFAMILY | SSF50156 | PDZ domain-like | IPR036034 | PDZ superfamily | 255 | 339 | 4.03E-9 |
| CDD | cd07560 | Peptidase_S41_CPP | IPR004447 | C-terminal-processing peptidase S41A | 381 | 553 | 6.37776E-72 |
| Pfam | PF00595 | PDZ domain | IPR001478 | PDZ domain | 257 | 337 | 1.2E-9 |
| Gene3D | G3DSA:3.30.750.44 | - | - | - | 188 | 531 | 8.6E-80 |
| SUPERFAMILY | SSF52096 | ClpP/crotonase | IPR029045 | ClpP/crotonase-like domain superfamily | 199 | 560 | 2.15E-69 |
| FunFam | G3DSA:3.30.750.44:FF:000007 | Periplasmic tail-specific protease | - | - | 187 | 273 | 2.7E-41 |
| CDD | cd00988 | PDZ_CTP_protease | - | - | 263 | 348 | 6.62996E-13 |
| SMART | SM00228 | pdz_new | IPR001478 | PDZ domain | 264 | 341 | 6.7E-5 |
| Gene3D | G3DSA:2.30.42.10 | - | IPR036034 | PDZ superfamily | 264 | 361 | 8.6E-80 |
| Coils | Coil | Coil | - | - | 206 | 226 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 654 | 687 | - |
| PANTHER | PTHR32060 | TAIL-SPECIFIC PROTEASE | - | - | 168 | 579 | 2.1E-75 |
| NCBIfam | TIGR00225 | JCVI: C-terminal processing peptidase | IPR004447 | C-terminal-processing peptidase S41A | 221 | 576 | 1.2E-95 |
| Coils | Coil | Coil | - | - | 617 | 651 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.