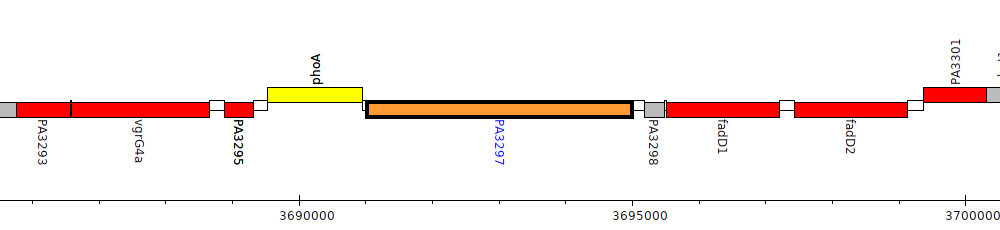
Pseudomonas aeruginosa PAO1, PA3297
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
27014238 | Reviewed by curator |
| Biological Process | GO:0006364 | rRNA processing | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
27014238 | Reviewed by curator |
| Molecular Function | GO:0004386 | helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00847
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003724 | RNA helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01967
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF07717 | Oligonucleotide/oligosaccharide-binding (OB)-fold | IPR011709 | DEAD-box helicase, OB fold | 647 | 723 | 8.0E-22 |
| FunFam | G3DSA:3.40.50.300:FF:000575 | ATP-dependent helicase hrpA | - | - | 27 | 247 | 4.6E-116 |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 87 | 391 | 9.2E-4 |
| CDD | cd18791 | SF2_C_RHA | - | - | 249 | 419 | 2.66818E-64 |
| Pfam | PF11898 | Domain of unknown function (DUF3418) | IPR024590 | RNA helicase HrpA, C-terminal | 736 | 1323 | 0.0 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 248 | 427 | 5.1E-57 |
| SMART | SM00847 | ha2_5 | IPR007502 | Helicase-associated domain | 471 | 561 | 5.9E-37 |
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 294 | 410 | 2.0E-11 |
| Gene3D | G3DSA:1.20.120.1080 | - | - | - | 456 | 552 | 3.0E-26 |
| SMART | SM00490 | helicmild6 | IPR001650 | Helicase, C-terminal | 311 | 411 | 1.6E-15 |
| FunFam | G3DSA:1.10.10.2130:FF:000001 | Pre-mRNA-splicing factor ATP-dependent RNA helicase | - | - | 433 | 476 | 6.3E-15 |
| Pfam | PF04408 | Helicase associated domain (HA2) | IPR007502 | Helicase-associated domain | 474 | 558 | 1.9E-18 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 74 | 591 | 2.14E-104 |
| PANTHER | PTHR18934 | ATP-DEPENDENT RNA HELICASE | - | - | 69 | 729 | 0.0 |
| FunFam | G3DSA:1.20.120.1080:FF:000005 | ATP-dependent helicase HrpA | - | - | 457 | 554 | 4.2E-40 |
| SMART | SM00487 | ultradead3 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 69 | 254 | 3.7E-24 |
| FunFam | G3DSA:3.40.50.300:FF:000439 | ATP-dependent RNA helicase HrpA | - | - | 248 | 427 | 2.1E-85 |
| CDD | cd17989 | DEXHc_HrpA | - | - | 72 | 244 | 2.64957E-112 |
| NCBIfam | TIGR01967 | JCVI: ATP-dependent RNA helicase HrpA | IPR010222 | RNA helicase HrpA | 13 | 1323 | 0.0 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 20 | 247 | 1.3E-76 |
| Coils | Coil | Coil | - | - | 786 | 806 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.