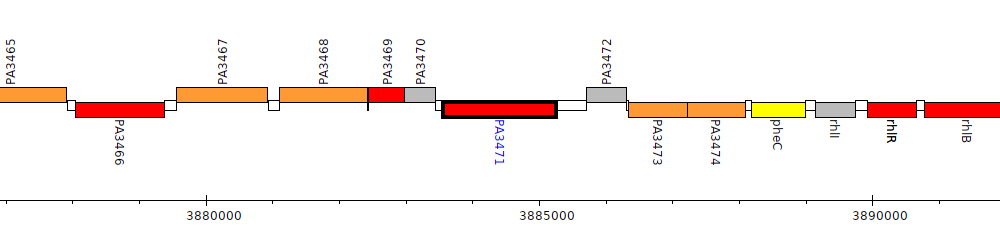
Pseudomonas aeruginosa PAO1, PA3471
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004470 | malic enzyme activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00919
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004471 | malate dehydrogenase (decarboxylating) (NAD+) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00620 | Pyruvate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 407 | 423 | 2.9E-72 |
| SMART | SM00919 | Malic_M_2 | IPR012302 | Malic enzyme, NAD-binding | 269 | 530 | 6.2E-129 |
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 145 | 174 | 2.9E-72 |
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 85 | 109 | 2.9E-72 |
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 181 | 203 | 2.9E-72 |
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 240 | 258 | 2.9E-72 |
| Gene3D | G3DSA:3.40.50.10380 | - | IPR037062 | Malic enzyme, N-terminal domain superfamily | 15 | 266 | 3.7E-110 |
| PANTHER | PTHR23406 | MALIC ENZYME-RELATED | - | - | 10 | 559 | 0.0 |
| SUPERFAMILY | SSF53223 | Aminoacid dehydrogenase-like, N-terminal domain | IPR046346 | Aminoacid dehydrogenase-like, N-terminal domain superfamily | 5 | 268 | 7.19E-107 |
| Pfam | PF00390 | Malic enzyme, N-terminal domain | IPR012301 | Malic enzyme, N-terminal domain | 79 | 259 | 3.8E-77 |
| Pfam | PF03949 | Malic enzyme, NAD binding domain | IPR012302 | Malic enzyme, NAD-binding | 269 | 529 | 2.9E-105 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 267 | 564 | 2.4E-105 |
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 296 | 312 | 2.9E-72 |
| SMART | SM01274 | malic_2 | IPR012301 | Malic enzyme, N-terminal domain | 79 | 259 | 9.1E-105 |
| PRINTS | PR00072 | Malic enzyme signature | IPR001891 | Malic oxidoreductase | 265 | 281 | 2.9E-72 |
| FunFam | G3DSA:3.40.50.10380:FF:000001 | NAD-dependent malic enzyme | - | - | 14 | 266 | 0.0 |
| FunFam | G3DSA:3.40.50.720:FF:000055 | NAD-dependent malic enzyme | - | - | 267 | 563 | 0.0 |
| PIRSF | PIRSF000106 | ME | IPR001891 | Malic oxidoreductase | 1 | 562 | 0.0 |
| Hamap | MF_01619 | NAD-dependent malic enzyme [maeA]. | IPR023667 | NAD-dependent malic enzyme, proteobacteria | 2 | 564 | 70.344315 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 269 | 561 | 1.0E-92 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.