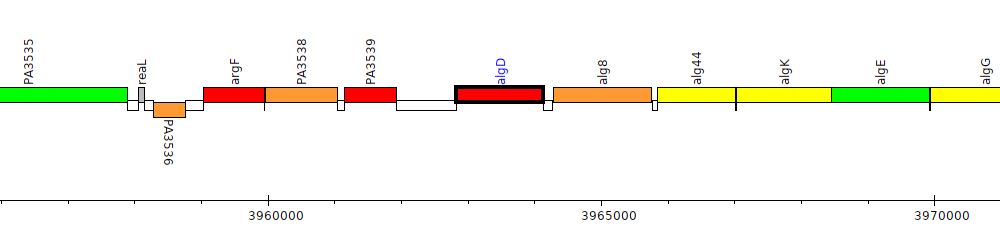
Pseudomonas aeruginosa PAO1, PA3540 (algD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006970 | response to osmotic stress | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
2496102 | Reviewed by curator |
| Molecular Function | GO:0047919 | GDP-mannose 6-dehydrogenase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
7521247 | Reviewed by curator |
| Biological Process | GO:0036460 | cellular response to cell envelope stress | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
17020580 | Reviewed by curator |
| Molecular Function | GO:0047919 | GDP-mannose 6-dehydrogenase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
2470755 | Reviewed by curator |
| Biological Process | GO:0009628 | response to abiotic stimulus | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
24496793 | Reviewed by curator |
| Biological Process | GO:0042121 | alginic acid biosynthetic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
7521247 | Reviewed by curator |
| Biological Process | GO:0050896 | response to stimulus | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0042121 | alginic acid biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
7521247 | Reviewed by curator |
| Biological Process | GO:0044010 | single-species biofilm formation | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
7521247 | Reviewed by curator |
| Biological Process | GO:0009607 | response to biotic stimulus | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
18455339 | Reviewed by curator |
| Molecular Function | GO:0051287 | NAD binding | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
12705829 | Reviewed by curator |
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | |
|||
| Biological Process | GO:0042121 | alginic acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF500135
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0047919 | GDP-mannose 6-dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF500135
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00984
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03026
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Alginate biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PWY-6082 | alginate biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Fructose and mannose metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00051 | Fructose and mannose metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00984 | UDP-glucose/GDP-mannose dehydrogenase family, central domain | IPR014026 | UDP-glucose/GDP-mannose dehydrogenase, dimerisation | 205 | 298 | 2.1E-28 |
| PANTHER | PTHR43750 | UDP-GLUCOSE 6-DEHYDROGENASE TUAD | - | - | 1 | 404 | 3.1E-114 |
| Pfam | PF03721 | UDP-glucose/GDP-mannose dehydrogenase family, NAD binding domain | IPR001732 | UDP-glucose/GDP-mannose dehydrogenase, N-terminal | 1 | 190 | 6.0E-63 |
| Pfam | PF03720 | UDP-glucose/GDP-mannose dehydrogenase family, UDP binding domain | IPR014027 | UDP-glucose/GDP-mannose dehydrogenase, C-terminal | 318 | 405 | 2.6E-12 |
| PIRSF | PIRSF000124 | UDPglc_GDPman_dh | IPR017476 | UDP-glucose/GDP-mannose dehydrogenase | 1 | 436 | 0.0 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 1 | 195 | 2.91E-51 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 1 | 202 | 5.9E-80 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 204 | 300 | 7.48E-28 |
| FunFam | G3DSA:3.40.50.720:FF:000573 | GDP-mannose 6-dehydrogenase | - | - | 1 | 202 | 0.0 |
| Gene3D | G3DSA:1.20.5.170 | - | - | - | 203 | 234 | 6.4E-20 |
| FunFam | G3DSA:3.40.50.720:FF:000664 | GDP-mannose 6-dehydrogenase | - | - | 235 | 436 | 0.0 |
| NCBIfam | TIGR03026 | JCVI: nucleotide sugar dehydrogenase | IPR017476 | UDP-glucose/GDP-mannose dehydrogenase | 1 | 410 | 1.1E-130 |
| SMART | SM00984 | UDPG_MGDP_dh_C_a_2_a | IPR014027 | UDP-glucose/GDP-mannose dehydrogenase, C-terminal | 317 | 425 | 2.2E-19 |
| SUPERFAMILY | SSF52413 | UDP-glucose/GDP-mannose dehydrogenase C-terminal domain | IPR036220 | UDP-glucose/GDP-mannose dehydrogenase, C-terminal domain superfamily | 305 | 434 | 2.62E-25 |
| PIRSF | PIRSF500135 | GDPman_DH | IPR028358 | GDP-mannose 6-dehydrogenase | 1 | 436 | 0.0 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 235 | 436 | 3.3E-58 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.