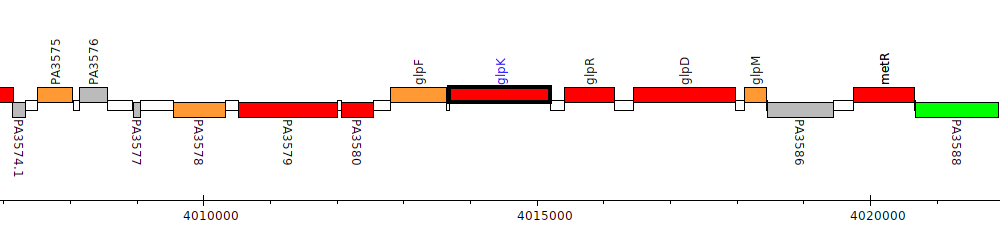
Pseudomonas aeruginosa PAO1, PA3582 (glpK)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0046486 | glycerolipid metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0008152 | metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004315 | 3-oxoacyl-[acyl-carrier-protein] synthase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004370 | glycerol kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00186
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000538
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006072 | glycerol-3-phosphate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00186
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016301 | kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000538
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | AERO-GLYCEROL-FERM-PWY | aerobic glycerol degradation I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00561 | Glycerolipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glycerol degradation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Glycerolipid metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02782 | FGGY family of carbohydrate kinases, C-terminal domain | IPR018485 | Carbohydrate kinase, FGGY, C-terminal | 265 | 455 | 1.2E-55 |
| SUPERFAMILY | SSF53067 | Actin-like ATPase domain | IPR043129 | ATPase, nucleotide binding domain | 258 | 500 | 1.51E-85 |
| SUPERFAMILY | SSF53067 | Actin-like ATPase domain | IPR043129 | ATPase, nucleotide binding domain | 8 | 259 | 1.68E-96 |
| Gene3D | G3DSA:3.30.420.40 | - | - | - | 258 | 503 | 1.2E-100 |
| FunFam | G3DSA:3.30.420.40:FF:000008 | Glycerol kinase | - | - | 5 | 257 | 0.0 |
| CDD | cd07786 | FGGY_EcGK_like | - | - | 9 | 495 | 0.0 |
| PANTHER | PTHR10196 | SUGAR KINASE | - | - | 10 | 500 | 0.0 |
| Gene3D | G3DSA:3.30.420.40 | - | - | - | 4 | 257 | 9.5E-100 |
| Hamap | MF_00186 | Glycerol kinase [glpK]. | IPR005999 | Glycerol kinase | 7 | 501 | 59.325039 |
| NCBIfam | TIGR01311 | JCVI: glycerol kinase GlpK | IPR005999 | Glycerol kinase | 8 | 500 | 0.0 |
| PIRSF | PIRSF000538 | GlpK | IPR000577 | Carbohydrate kinase, FGGY | 7 | 504 | 0.0 |
| Pfam | PF00370 | FGGY family of carbohydrate kinases, N-terminal domain | IPR018484 | Carbohydrate kinase, FGGY, N-terminal | 9 | 256 | 4.6E-88 |
| FunFam | G3DSA:3.30.420.40:FF:000007 | Glycerol kinase | - | - | 258 | 500 | 2.4E-119 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.