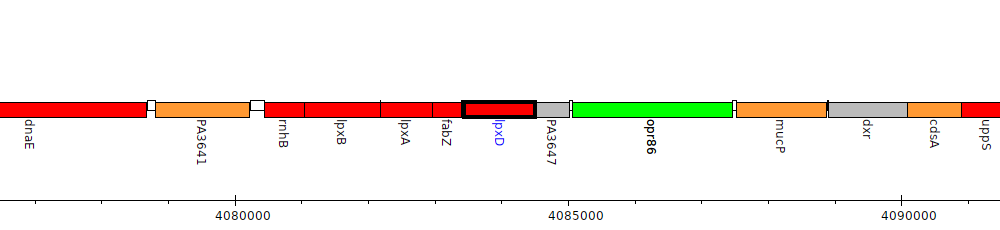
Pseudomonas aeruginosa PAO1, PA3646 (lpxD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009245 | lipid A biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA14_17180
|
ECO:0000250 sequence similarity evidence used in manual assertion |
25173672 | Reviewed by curator |
| Molecular Function | GO:0016746 | transferase activity, transferring acyl groups | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8293817 | Reviewed by curator |
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0009245 | lipid A biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43378
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016747 | transferase activity, transferring acyl groups other than amino-acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF04613
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016410 | N-acyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43378
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00540 | Lipopolysaccharide biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Lipopolysaccharide biosynthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | Coil | - | - | 319 | 346 | - |
| Gene3D | G3DSA:1.20.5.170 | - | - | - | 313 | 353 | 4.4E-20 |
| Gene3D | G3DSA:3.40.1390.10 | - | - | - | 2 | 101 | 2.2E-37 |
| PANTHER | PTHR43378 | UDP-3-O-ACYLGLUCOSAMINE N-ACYLTRANSFERASE | IPR007691 | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase LpxD | 7 | 340 | 1.8E-76 |
| NCBIfam | TIGR01853 | JCVI: UDP-3-O-(3-hydroxymyristoyl)glucosamine N-acyltransferase | IPR007691 | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase LpxD | 11 | 332 | 2.3E-117 |
| Hamap | MF_00523 | UDP-3-O-(3-hydroxymyristoyl)glucosamine N-acyltransferase [lpxD]. | IPR007691 | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase LpxD | 12 | 329 | 36.371384 |
| Pfam | PF00132 | Bacterial transferase hexapeptide (six repeats) | IPR001451 | Hexapeptide repeat | 149 | 183 | 1.5E-6 |
| CDD | cd03352 | LbH_LpxD | IPR007691 | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase LpxD | 112 | 317 | 9.22551E-113 |
| Pfam | PF04613 | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase, LpxD | IPR020573 | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase, non-repeat region | 25 | 92 | 8.7E-22 |
| SUPERFAMILY | SSF51161 | Trimeric LpxA-like enzymes | IPR011004 | Trimeric LpxA-like superfamily | 35 | 320 | 2.51E-80 |
| Gene3D | G3DSA:2.160.10.10 | Hexapeptide repeat proteins | - | - | 102 | 312 | 1.6E-68 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.