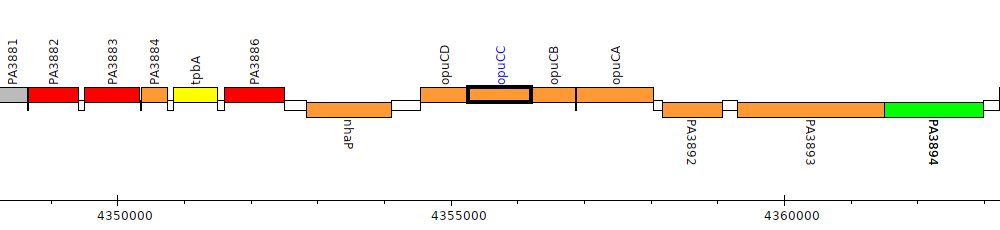
Pseudomonas aeruginosa PAO1, PA3889 (opuCC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0015838 | amino-acid betaine transport |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:Q87WH3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17660277 | Reviewed by curator |
| Molecular Function | GO:0015226 | carnitine transmembrane transporter activity |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:Q87WH3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17660277 | Reviewed by curator |
| Biological Process | GO:0071475 | cellular hyperosmotic salinity response |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:Q87WH3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17660277 | Reviewed by curator |
| Biological Process | GO:0031460 | glycine betaine transport |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:Q87WH3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17660277 | Reviewed by curator |
| Cellular Component | GO:0043190 | ATP-binding cassette (ABC) transporter complex |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:Q87WH3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17660277 | Reviewed by curator |
| Cellular Component | GO:0042597 | periplasmic space |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:Q87WH3
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17660277 | Reviewed by curator |
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04069
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0043190 | ATP-binding cassette (ABC) transporter complex |
Inferred from Sequence Model
Term mapped from: InterPro:PF04069
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF04069
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae02010 | ABC transporters | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR30006 | THIAMINE-BINDING PERIPLASMIC PROTEIN-RELATED | - | - | 1 | 278 | 4.3E-16 |
| Gene3D | G3DSA:3.40.190.120 | Osmoprotection protein (prox); domain 2 | - | - | 124 | 238 | 1.5E-91 |
| SUPERFAMILY | SSF53850 | Periplasmic binding protein-like II | - | - | 25 | 299 | 2.23E-69 |
| CDD | cd13611 | PBP2_YehZ | - | - | 25 | 295 | 0.0 |
| Gene3D | G3DSA:3.40.190.10 | - | - | - | 28 | 297 | 1.5E-91 |
| Pfam | PF04069 | Substrate binding domain of ABC-type glycine betaine transport system | IPR007210 | ABC-type glycine betaine transport system, substrate-binding domain | 25 | 296 | 9.0E-69 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.