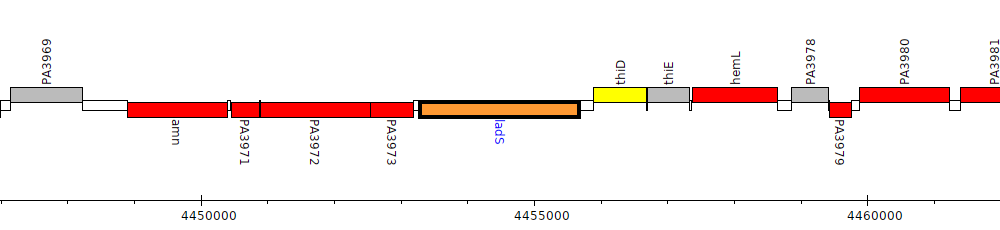
Pseudomonas aeruginosa PAO1, PA3974 (ladS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:cd00082
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00082
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:SM00448
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Two-component System |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52172 | CheY-like | IPR011006 | CheY-like superfamily | 667 | 785 | 1.05E-39 |
| Gene3D | G3DSA:1.10.287.130 | - | - | - | 383 | 481 | 2.4E-19 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 588 | 598 | 1.0E-12 |
| SMART | SM00388 | HisKA_10 | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 418 | 483 | 5.3E-16 |
| Pfam | PF07696 | 7TMR-DISM extracellular 2 | IPR011622 | 7TM-DISM receptor, extracellular domain, type 2 | 32 | 171 | 5.4E-37 |
| Gene3D | G3DSA:3.40.50.2300 | - | - | - | 669 | 791 | 2.5E-47 |
| Gene3D | G3DSA:3.30.565.10 | - | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 482 | 647 | 4.0E-42 |
| FunFam | G3DSA:3.30.565.10:FF:000010 | Sensor histidine kinase RcsC | - | - | 482 | 647 | 7.0E-45 |
| SUPERFAMILY | SSF47384 | Homodimeric domain of signal transducing histidine kinase | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 404 | 484 | 1.12E-16 |
| Gene3D | G3DSA:2.60.40.2380 | - | - | - | 29 | 178 | 3.6E-38 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 629 | 642 | 1.0E-12 |
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 670 | 780 | 4.7E-27 |
| CDD | cd17546 | REC_hyHK_CKI1_RcsC-like | - | - | 670 | 781 | 8.55386E-50 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 570 | 584 | 1.0E-12 |
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 530 | 644 | 5.1E-23 |
| Pfam | PF07695 | 7TM diverse intracellular signalling | IPR011623 | 7TM-DISM receptor, extracellular domain, type 1 | 184 | 386 | 5.5E-56 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 605 | 623 | 1.0E-12 |
| Coils | Coil | Coil | - | - | 391 | 425 | - |
| SMART | SM00448 | REC_2 | IPR001789 | Signal transduction response regulator, receiver domain | 668 | 781 | 5.5E-39 |
| PANTHER | PTHR45339 | HYBRID SIGNAL TRANSDUCTION HISTIDINE KINASE J | - | - | 365 | 649 | 3.2E-121 |
| CDD | cd00082 | HisKA | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 416 | 477 | 8.6091E-13 |
| SUPERFAMILY | SSF55874 | ATPase domain of HSP90 chaperone/DNA topoisomerase II/histidine kinase | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 472 | 642 | 5.76E-37 |
| CDD | cd16922 | HATPase_EvgS-ArcB-TorS-like | - | - | 535 | 643 | 5.8109E-49 |
| SMART | SM00387 | HKATPase_4 | IPR003594 | Histidine kinase/HSP90-like ATPase | 530 | 645 | 1.2E-29 |
| Pfam | PF00512 | His Kinase A (phospho-acceptor) domain | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 419 | 482 | 2.2E-14 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.