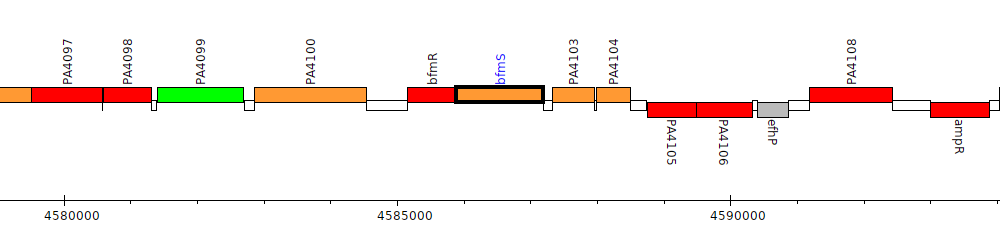
Pseudomonas aeruginosa PAO1, PA4102 (bfmS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009405 | pathogenesis | Inferred from Mutant Phenotype | ECO:0000016 loss-of-function mutant phenotype evidence |
25166864 | Reviewed by curator |
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
19936057 | Reviewed by curator |
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF00672
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:PF00672
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00388
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Two-component System |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 329 | 433 | 2.1E-21 |
| SMART | SM00388 | HisKA_10 | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 228 | 286 | 0.0079 |
| SMART | SM00387 | HKATPase_4 | IPR003594 | Histidine kinase/HSP90-like ATPase | 326 | 434 | 1.3E-29 |
| Gene3D | G3DSA:1.10.287.130 | - | - | - | 172 | 280 | 7.0E-15 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 379 | 389 | 2.9E-10 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 361 | 375 | 2.9E-10 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 396 | 414 | 2.9E-10 |
| CDD | cd00082 | HisKA | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 226 | 276 | 0.00382815 |
| SMART | SM00304 | HAMP_11 | IPR003660 | HAMP domain | 175 | 227 | 1.4E-5 |
| PANTHER | PTHR44936 | SENSOR PROTEIN CREC | - | - | 7 | 434 | 6.5E-65 |
| SUPERFAMILY | SSF55874 | ATPase domain of HSP90 chaperone/DNA topoisomerase II/histidine kinase | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 292 | 432 | 2.36E-32 |
| Pfam | PF00672 | HAMP domain | IPR003660 | HAMP domain | 173 | 222 | 3.9E-6 |
| FunFam | G3DSA:3.30.565.10:FF:000253 | Two-component sensor histidine kinase | - | - | 287 | 433 | 5.6E-97 |
| SUPERFAMILY | SSF47384 | Homodimeric domain of signal transducing histidine kinase | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 212 | 282 | 4.71E-9 |
| Gene3D | G3DSA:3.30.565.10 | - | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 287 | 433 | 6.2E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.