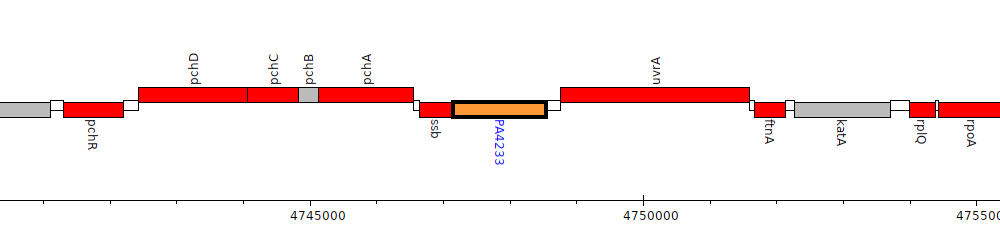
Pseudomonas aeruginosa PAO1, PA4233
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 394 | 457 | 1.5E-25 |
| CDD | cd17472 | MFS_YajR_like | - | - | 20 | 393 | 0.0 |
| Pfam | PF07690 | Major Facilitator Superfamily | IPR011701 | Major facilitator superfamily | 21 | 358 | 4.3E-46 |
| PANTHER | PTHR23517 | RESISTANCE PROTEIN MDTM, PUTATIVE-RELATED-RELATED | - | - | 12 | 392 | 1.7E-53 |
| SUPERFAMILY | SSF103473 | MFS general substrate transporter | IPR036259 | MFS transporter superfamily | 14 | 395 | 3.4E-69 |
| Gene3D | G3DSA:1.20.1250.20 | MFS general substrate transporter like domains | IPR036259 | MFS transporter superfamily | 7 | 393 | 2.1E-96 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.