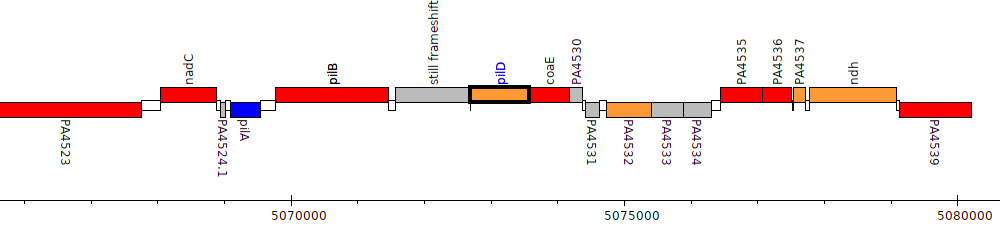
Pseudomonas aeruginosa PAO1, PA4528 (pilD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006480 | N-terminal protein amino acid methylation | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
23255525 | Reviewed by curator |
| Cellular Component | GO:0005886 | plasma membrane | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9466269 | Reviewed by curator |
| Molecular Function | GO:0004190 | aspartic-type endopeptidase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
23255525 | Reviewed by curator |
| Molecular Function | GO:0008170 | N-methyltransferase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8096341 | Reviewed by curator |
| Biological Process | GO:0006465 | signal peptide processing | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8096341 | Reviewed by curator |
| Biological Process | GO:0043683 | type IV pilus biogenesis | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
1671384 | Reviewed by curator |
| Cellular Component | GO:0015627 | type II protein secretion system complex | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
1588814 | Reviewed by curator |
| Biological Process | GO:0009297 | pilus assembly | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9668097 | Reviewed by curator |
| Molecular Function | GO:0004190 | aspartic-type endopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01478
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF01478
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Pilin biosynthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF06750 | Bacterial Peptidase A24 N-terminal domain | IPR010627 | Peptidase A24A, N-terminal | 20 | 126 | 7.0E-34 |
| PANTHER | PTHR30487 | TYPE 4 PREPILIN-LIKE PROTEINS LEADER PEPTIDE-PROCESSING ENZYME | - | - | 13 | 286 | 8.4E-89 |
| PRINTS | PR00864 | Type IV prepilin cysteine protease (C20) family signature | IPR014032 | Peptidase A24A, prepilin type IV, bacterial | 145 | 162 | 1.5E-26 |
| PRINTS | PR00864 | Type IV prepilin cysteine protease (C20) family signature | IPR014032 | Peptidase A24A, prepilin type IV, bacterial | 213 | 224 | 1.5E-26 |
| Pfam | PF01478 | Type IV leader peptidase family | IPR000045 | Prepilin type IV endopeptidase, peptidase domain | 137 | 245 | 8.9E-25 |
| FunFam | G3DSA:1.20.120.1220:FF:000001 | Type 4 prepilin-like proteins leader peptide-processing enzyme | - | - | 128 | 286 | 9.7E-76 |
| PRINTS | PR00864 | Type IV prepilin cysteine protease (C20) family signature | IPR014032 | Peptidase A24A, prepilin type IV, bacterial | 202 | 212 | 1.5E-26 |
| PRINTS | PR00864 | Type IV prepilin cysteine protease (C20) family signature | IPR014032 | Peptidase A24A, prepilin type IV, bacterial | 225 | 240 | 1.5E-26 |
| Gene3D | G3DSA:1.20.120.1220 | - | - | - | 111 | 286 | 2.8E-13 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.