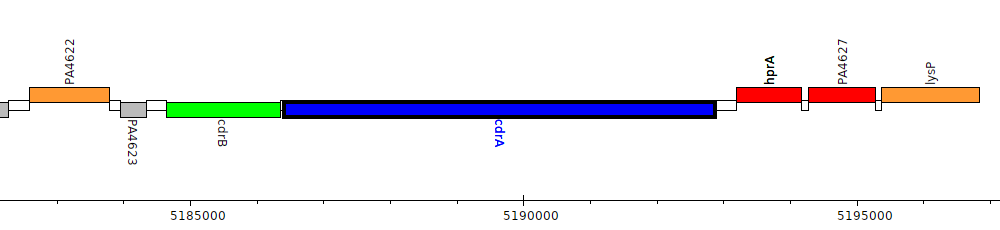
Pseudomonas aeruginosa PAO1, PA4625 (cdrA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0097312 | obsolete bacterial biofilm matrix component | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20088866 | Reviewed by curator |
| Cellular Component | GO:0098046 | type V protein secretion system complex |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P12255
|
ECO:0000250 sequence similarity evidence used in manual assertion |
20088866 | Reviewed by curator |
| Biological Process | GO:0044010 | single-species biofilm formation | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20088866 | Reviewed by curator |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Two-partner secretion system |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1211 | 1289 | 5.4E-5 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1372 | 1450 | 5.4E-5 |
| NCBIfam | TIGR01901 | JCVI: filamentous hemagglutinin family N-terminal domain | IPR008638 | Filamentous haemagglutinin FhaB/tRNA nuclease CdiA-like, TPS domain | 74 | 149 | 2.3E-18 |
| PANTHER | PTHR12338 | AUTOTRANSPORTER | - | - | 1691 | 1857 | 0.0 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1476 | 1551 | 1.5E-20 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1453 | 1525 | 2.7E-5 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1718 | 1793 | 3.4E-20 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 893 | 954 | 4.9E-5 |
| FunFam | G3DSA:2.160.20.10:FF:000109 | Hemagglutination activity domain protein | - | - | 42 | 437 | 0.0 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1154 | 1228 | 1.5E-19 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1234 | 1309 | 3.4E-20 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 796 | 886 | 5.1E-7 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1776 | 1847 | 2.8E-5 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1131 | 1197 | 3.3E-5 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1557 | 1632 | 3.4E-20 |
| SMART | SM00912 | Haemagg_act_2a | IPR008638 | Filamentous haemagglutinin FhaB/tRNA nuclease CdiA-like, TPS domain | 40 | 152 | 1.2E-35 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 992 | 1067 | 1.9E-19 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1695 | 1773 | 5.4E-5 |
| SUPERFAMILY | SSF51126 | Pectin lyase-like | IPR011050 | Pectin lyase fold/virulence factor | 63 | 336 | 1.48E-43 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1880 | 1954 | 3.6E-19 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1960 | 2033 | 4.3E-16 |
| Gene3D | G3DSA:2.160.20.10 | - | IPR012334 | Pectin lyase fold | 42 | 424 | 5.3E-89 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1799 | 1874 | 2.8E-20 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1315 | 1389 | 3.2E-19 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1050 | 1121 | 3.7E-6 |
| Gene3D | G3DSA:3.30.160.710 | - | - | - | 1534 | 1612 | 5.4E-5 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 824 | 906 | 1.4E-11 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 912 | 985 | 7.8E-20 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1638 | 1712 | 3.2E-19 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1395 | 1470 | 3.4E-20 |
| Pfam | PF05860 | TPS secretion domain | IPR008638 | Filamentous haemagglutinin FhaB/tRNA nuclease CdiA-like, TPS domain | 40 | 303 | 6.1E-44 |
| Pfam | PF18676 | MBG domain (YGX type) | IPR041286 | MBG domain, YGX type | 1073 | 1148 | 1.1E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.