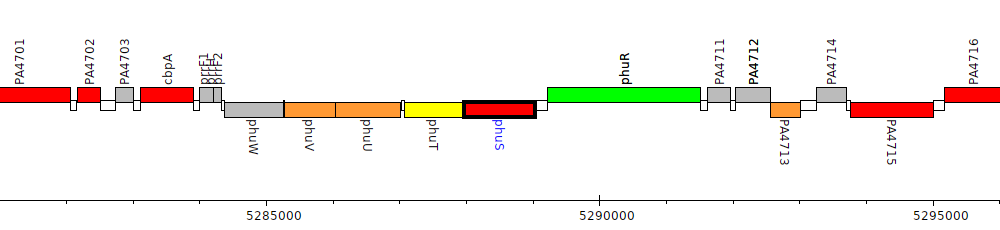
Pseudomonas aeruginosa PAO1, PA4709 (phuS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0020037 | heme binding | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
10658665 | Reviewed by curator |
| Biological Process | GO:0006826 | iron ion transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF05171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd16831 | HemS-like_C | - | - | 197 | 350 | 7.67853E-96 |
| Gene3D | G3DSA:3.40.1570.10 | HemS/ChuS/ChuX like domains | - | - | 18 | 294 | 0.0 |
| CDD | cd16830 | HemS-like_N | - | - | 14 | 167 | 1.81614E-82 |
| SUPERFAMILY | SSF144064 | Heme iron utilization protein-like | - | - | 14 | 351 | 2.94E-126 |
| Pfam | PF05171 | Haemin-degrading HemS.ChuX domain | IPR007845 | Haemin-degrading HemS/ChuX domain | 220 | 351 | 9.9E-43 |
| Pfam | PF05171 | Haemin-degrading HemS.ChuX domain | IPR007845 | Haemin-degrading HemS/ChuX domain | 38 | 167 | 6.6E-43 |
| Gene3D | G3DSA:3.40.1570.10 | HemS/ChuS/ChuX like domains | - | - | 91 | 346 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.