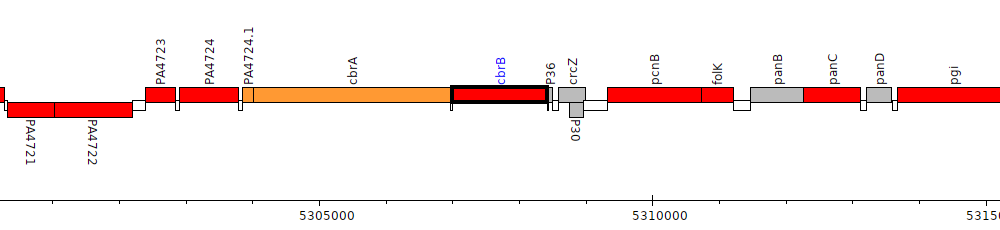
Pseudomonas aeruginosa PAO1, PA4726 (cbrB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044248 | cellular catabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:2000147 | positive regulation of cell motility | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
21169488 | Reviewed by curator |
| Molecular Function | GO:0000156 | phosphorelay response regulator activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11401699 | Reviewed by curator |
| Biological Process | GO:1900081 | regulation of arginine catabolic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
11401699 | Reviewed by curator |
| Biological Process | GO:2001158 | positive regulation of proline catabolic process to glutamate | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
11401699 | Reviewed by curator |
| Biological Process | GO:0090368 | regulation of ornithine metabolic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
11401699 | Reviewed by curator |
| Biological Process | GO:0015976 | carbon utilization | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00158
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008134 | transcription factor binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00158
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF00158
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:SM00448
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR32071 | TRANSCRIPTIONAL REGULATORY PROTEIN | - | - | 1 | 470 | 0.0 |
| CDD | cd00156 | REC | - | - | 5 | 99 | 7.79353E-28 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 148 | 390 | 1.74E-72 |
| Pfam | PF00158 | Sigma-54 interaction domain | IPR002078 | RNA polymerase sigma factor 54 interaction domain | 149 | 315 | 5.5E-73 |
| SUPERFAMILY | SSF52172 | CheY-like | IPR011006 | CheY-like superfamily | 1 | 194 | 2.99E-37 |
| Gene3D | G3DSA:3.40.50.2300 | - | - | - | 2 | 129 | 7.5E-29 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 141 | 319 | 2.9E-69 |
| SMART | SM00448 | REC_2 | IPR001789 | Signal transduction response regulator, receiver domain | 2 | 109 | 6.2E-28 |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 169 | 334 | 7.6E-11 |
| CDD | cd00009 | AAA | - | - | 156 | 313 | 5.10072E-28 |
| FunFam | G3DSA:3.40.50.300:FF:000006 | DNA-binding transcriptional regulator NtrC | - | - | 143 | 319 | 5.0E-80 |
| FunFam | G3DSA:3.40.50.2300:FF:000794 | - | - | - | 2 | 129 | 4.6E-85 |
| Gene3D | G3DSA:1.10.8.60 | - | - | - | 320 | 397 | 1.3E-21 |
| FunFam | G3DSA:1.10.8.60:FF:000120 | Sigma-54-dependent Fis family transcriptional regulator | - | - | 321 | 397 | 9.3E-30 |
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 4 | 109 | 2.2E-21 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.