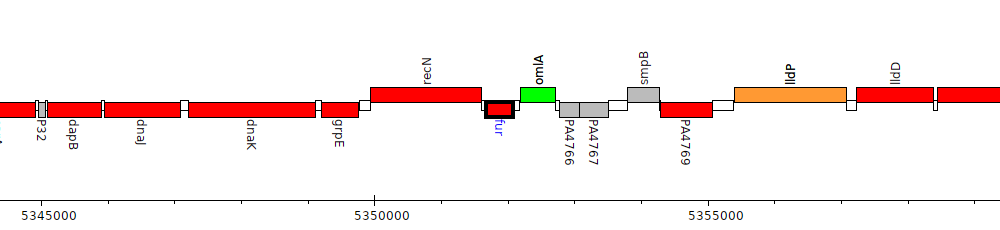
Pseudomonas aeruginosa PAO1, PA4764 (fur)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0008198 | ferrous iron binding | Traceable Author Statement | ECO:0000304 traceable author statement used in manual assertion |
8522528 | Reviewed by curator |
| Biological Process | GO:2000470 | positive regulation of peroxidase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
8763923 | Reviewed by curator |
| Biological Process | GO:1900705 | negative regulation of siderophore biosynthetic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
8885270 | Reviewed by curator |
| Biological Process | GO:0051350 | negative regulation of lyase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
9045799 | Reviewed by curator |
| Molecular Function | GO:0000976 | transcription regulatory region sequence-specific DNA binding | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8522528 | Reviewed by curator |
| Biological Process | GO:1901668 | regulation of superoxide dismutase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
8763923 | Reviewed by curator |
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd07153
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:cd07153
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF46785 | Winged helix DNA-binding domain | IPR036390 | Winged helix DNA-binding domain superfamily | 6 | 131 | 4.89E-41 |
| PANTHER | PTHR33202 | ZINC UPTAKE REGULATION PROTEIN | IPR002481 | Ferric-uptake regulator | 5 | 130 | 3.0E-33 |
| FunFam | G3DSA:3.30.1490.190:FF:000001 | Ferric uptake regulation protein | - | - | 84 | 134 | 1.3E-23 |
| FunFam | G3DSA:1.10.10.10:FF:000007 | Ferric uptake regulation protein | - | - | 1 | 83 | 3.3E-39 |
| Pfam | PF01475 | Ferric uptake regulator family | IPR002481 | Ferric-uptake regulator | 9 | 129 | 2.1E-46 |
| CDD | cd07153 | Fur_like | IPR002481 | Ferric-uptake regulator | 17 | 129 | 1.43511E-41 |
| Coils | Coil | Coil | - | - | 25 | 45 | - |
| Gene3D | G3DSA:3.30.1490.190 | - | IPR043135 | Ferric-uptake regulator, C-terminal domain | 84 | 134 | 4.5E-19 |
| Gene3D | G3DSA:1.10.10.10 | - | IPR036388 | Winged helix-like DNA-binding domain superfamily | 1 | 83 | 7.5E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.