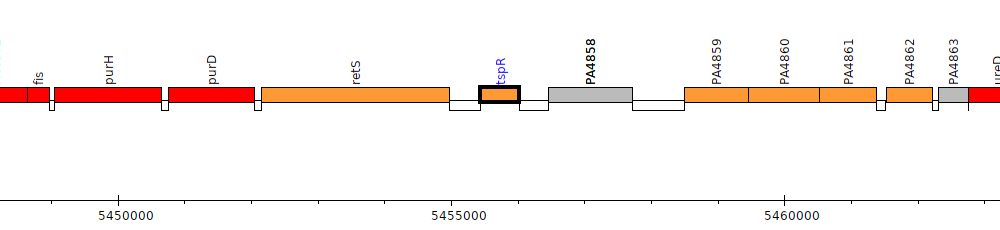
Pseudomonas aeruginosa PAO1, PA4857 (tspR)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0050714 | positive regulation of protein secretion | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
26858696 | Reviewed by curator |
| Biological Process | GO:0071978 | bacterial-type flagellum-dependent swarming motility | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
26858696 | Reviewed by curator |
| Biological Process | GO:1902021 | regulation of bacterial-type flagellum-dependent cell motility | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
26858696 | Reviewed by curator |
| Biological Process | GO:1900191 | negative regulation of single-species biofilm formation | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
26858696 | Reviewed by curator |
| Biological Process | GO:0030254 | protein secretion by the type III secretion system | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
26858696 | Reviewed by curator |
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR33508
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR33508 | UPF0056 MEMBRANE PROTEIN YHCE | IPR002771 | Multiple antibiotic resistance (MarC)-related | 2 | 193 | 5.9E-49 |
| NCBIfam | TIGR00427 | JCVI: NAAT family transporter | IPR002771 | Multiple antibiotic resistance (MarC)-related | 3 | 190 | 1.6E-31 |
| Pfam | PF01914 | MarC family integral membrane protein | IPR002771 | Multiple antibiotic resistance (MarC)-related | 2 | 193 | 7.7E-37 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.