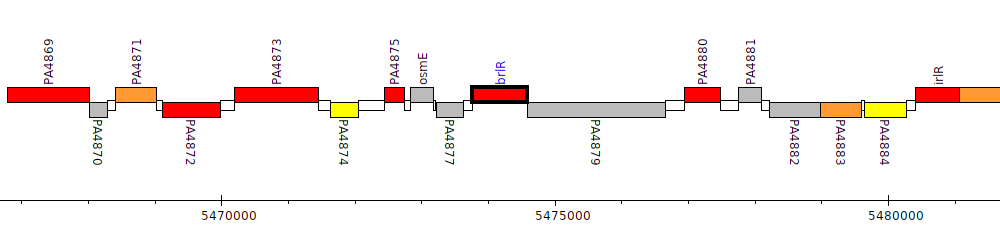
Pseudomonas aeruginosa PAO1, PA4878 (brlR)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0045893 | positive regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
22730129 | Reviewed by curator |
| Biological Process | GO:0045892 | negative regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
23935054 | Reviewed by curator |
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR30204
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00422
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:SM00422
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14526 | Integron-associated effector binding protein | IPR029441 | Integron-associated effector binding protein | 123 | 269 | 1.2E-36 |
| Gene3D | G3DSA:3.20.80.10 | Regulatory factor, effector binding domain | IPR011256 | Regulatory factor, effector binding domain superfamily | 124 | 270 | 7.2E-39 |
| SMART | SM00422 | merrmega3 | IPR000551 | MerR-type HTH domain | 2 | 72 | 4.8E-20 |
| FunFam | G3DSA:3.20.80.10:FF:000016 | Probable transcriptional regulator | - | - | 124 | 270 | 1.8E-103 |
| SMART | SM00871 | AraC_E_bind_2 | IPR010499 | Bacterial transcription activator, effector binding | 120 | 270 | 1.4E-33 |
| SUPERFAMILY | SSF55136 | Probable bacterial effector-binding domain | IPR011256 | Regulatory factor, effector binding domain superfamily | 120 | 269 | 1.05E-32 |
| CDD | cd01107 | HTH_BmrR | - | - | 2 | 112 | 1.83943E-42 |
| PANTHER | PTHR30204 | REDOX-CYCLING DRUG-SENSING TRANSCRIPTIONAL ACTIVATOR SOXR | IPR047057 | MerR transcriptional regulator | 1 | 112 | 2.4E-21 |
| Pfam | PF13411 | MerR HTH family regulatory protein | IPR000551 | MerR-type HTH domain | 3 | 68 | 1.0E-15 |
| SUPERFAMILY | SSF46955 | Putative DNA-binding domain | IPR009061 | Putative DNA-binding domain superfamily | 2 | 108 | 1.31E-25 |
| Gene3D | G3DSA:1.10.1660.10 | - | - | - | 1 | 123 | 3.4E-26 |
| FunFam | G3DSA:1.10.1660.10:FF:000016 | MerR family transcriptional regulator | - | - | 1 | 123 | 6.5E-61 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.