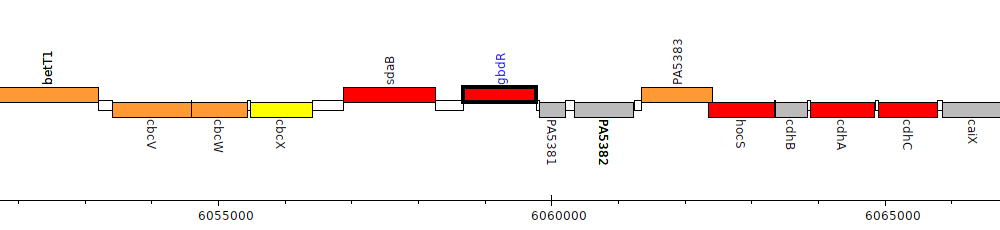
Pseudomonas aeruginosa PAO1, PA5380 (gbdR)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0031457 | glycine betaine catabolic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
17951379 | Reviewed by curator |
| Biological Process | GO:0010922 | positive regulation of phosphatase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
19103776 | Reviewed by curator |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
24097953 | Reviewed by curator |
| Biological Process | GO:0045764 | positive regulation of cellular amino acid metabolic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
17951379 | Reviewed by curator |
| Biological Process | GO:0045764 | positive regulation of cellular amino acid metabolic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
17951379 | Reviewed by curator |
| Biological Process | GO:0010922 | positive regulation of phosphatase activity | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
19103776 | Reviewed by curator |
| Biological Process | GO:0009896 | positive regulation of catabolic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
17951379 | Reviewed by curator |
| Biological Process | GO:0010518 | positive regulation of phospholipase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
19103776 | Reviewed by curator |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
17951379 | Reviewed by curator |
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:SM00342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00342
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | glycine betaine catabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00342 | aracneu4 | IPR018060 | DNA binding HTH domain, AraC-type | 240 | 323 | 9.5E-29 |
| FunFam | G3DSA:1.10.10.60:FF:000090 | Transcriptional regulator ArgR, AraC family | - | - | 220 | 325 | 2.2E-58 |
| SUPERFAMILY | SSF46689 | Homeodomain-like | IPR009057 | Homeobox-like domain superfamily | 224 | 274 | 2.15E-9 |
| Pfam | PF12833 | Helix-turn-helix domain | IPR018060 | DNA binding HTH domain, AraC-type | 246 | 324 | 1.2E-22 |
| Gene3D | G3DSA:1.10.10.60 | - | - | - | 220 | 326 | 4.7E-30 |
| SUPERFAMILY | SSF52317 | Class I glutamine amidotransferase-like | IPR029062 | Class I glutamine amidotransferase-like | 15 | 205 | 4.07E-48 |
| SUPERFAMILY | SSF46689 | Homeodomain-like | IPR009057 | Homeobox-like domain superfamily | 276 | 325 | 8.06E-12 |
| Pfam | PF01965 | DJ-1/PfpI family | IPR002818 | DJ-1/PfpI | 20 | 183 | 2.8E-17 |
| PANTHER | PTHR43130 | ARAC-FAMILY TRANSCRIPTIONAL REGULATOR | - | - | 13 | 263 | 5.8E-49 |
| FunFam | G3DSA:3.40.50.880:FF:000061 | AraC family transcriptional regulator | - | - | 12 | 213 | 0.0 |
| CDD | cd03136 | GATase1_AraC_ArgR_like | - | - | 18 | 202 | 5.86732E-97 |
| Gene3D | G3DSA:3.40.50.880 | - | IPR029062 | Class I glutamine amidotransferase-like | 12 | 213 | 5.1E-58 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.