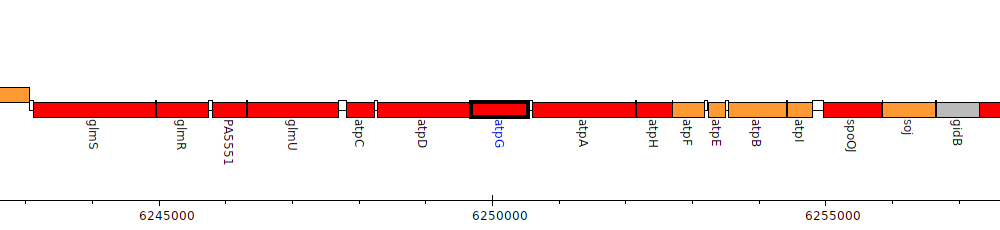
Pseudomonas aeruginosa PAO1, PA5555 (atpG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046961 | proton-transporting ATPase activity, rotational mechanism | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
146491 | Reviewed by curator |
| Molecular Function | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
146491 | Reviewed by curator |
| Cellular Component | GO:0045261 | proton-transporting ATP synthase complex, catalytic core F(1) | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2863270 | Reviewed by curator |
| Biological Process | GO:0015986 | ATP synthesis coupled proton transport |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11693
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11693
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0045261 | proton-transporting ATP synthase complex, catalytic core F(1) |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11693
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Oxidative phosphorylation |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-7219 | adenosine ribonucleotides de novo biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00190 | Oxidative phosphorylation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:1.10.287.80:FF:000005 | ATP synthase gamma chain | - | - | 196 | 286 | 1.3E-35 |
| PRINTS | PR00126 | ATP synthase gamma subunit signature | IPR000131 | ATP synthase, F1 complex, gamma subunit | 73 | 92 | 4.4E-37 |
| FunFam | G3DSA:3.40.1380.10:FF:000001 | ATP synthase gamma chain | - | - | 27 | 245 | 4.6E-122 |
| SUPERFAMILY | SSF52943 | ATP synthase (F1-ATPase), gamma subunit | IPR035968 | ATP synthase, F1 complex, gamma subunit superfamily | 2 | 286 | 1.83E-95 |
| PRINTS | PR00126 | ATP synthase gamma subunit signature | IPR000131 | ATP synthase, F1 complex, gamma subunit | 264 | 285 | 4.4E-37 |
| Gene3D | G3DSA:3.40.1380.10 | - | - | - | 27 | 245 | 3.0E-98 |
| Pfam | PF00231 | ATP synthase | IPR000131 | ATP synthase, F1 complex, gamma subunit | 5 | 285 | 1.9E-95 |
| NCBIfam | TIGR01146 | JCVI: ATP synthase F1 subunit gamma | IPR000131 | ATP synthase, F1 complex, gamma subunit | 1 | 286 | 3.8E-106 |
| CDD | cd12151 | F1-ATPase_gamma | IPR000131 | ATP synthase, F1 complex, gamma subunit | 2 | 283 | 1.3686E-120 |
| PRINTS | PR00126 | ATP synthase gamma subunit signature | IPR000131 | ATP synthase, F1 complex, gamma subunit | 166 | 183 | 4.4E-37 |
| PANTHER | PTHR11693 | ATP SYNTHASE GAMMA CHAIN | IPR000131 | ATP synthase, F1 complex, gamma subunit | 2 | 285 | 1.1E-54 |
| Gene3D | G3DSA:1.10.287.80 | ATP synthase, gamma subunit, helix hairpin domain | - | - | 5 | 283 | 3.0E-98 |
| Hamap | MF_00815 | ATP synthase gamma chain [atpG]. | IPR000131 | ATP synthase, F1 complex, gamma subunit | 1 | 286 | 36.71965 |
| PRINTS | PR00126 | ATP synthase gamma subunit signature | IPR000131 | ATP synthase, F1 complex, gamma subunit | 233 | 252 | 4.4E-37 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.