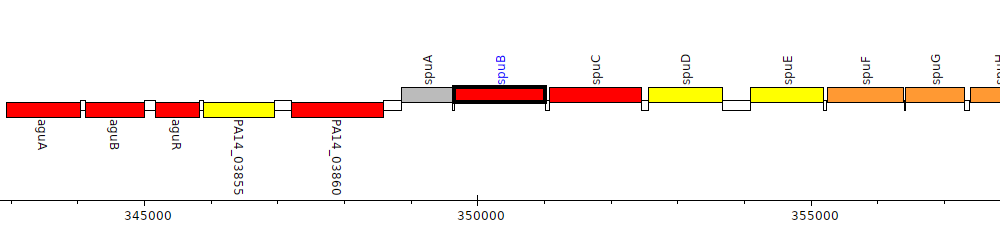
Pseudomonas aeruginosa UCBPP-PA14, PA14_03880 (spuB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0046203 | spermidine catabolic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0298
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
21622750 | |
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55931
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004356 | glutamate-ammonia ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.10.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006807 | nitrogen compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.10.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006542 | glutamine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.10.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00220 | Arginine biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00910 | Nitrogen metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00250 | Alanine, aspartate and glutamate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM01230 | Gln_synt_C_2 | IPR008146 | Glutamine synthetase, catalytic domain | 114 | 370 | 2.5E-112 |
| SUPERFAMILY | SSF55931 | Glutamine synthetase/guanido kinase | IPR014746 | Glutamine synthetase/guanido kinase, catalytic domain | 114 | 449 | 2.35E-106 |
| FunFam | G3DSA:3.10.20.70:FF:000011 | Glutamine synthetase | - | - | 4 | 115 | 1.1E-61 |
| Gene3D | G3DSA:3.10.20.70 | - | IPR036651 | Glutamine synthetase, N-terminal domain superfamily | 3 | 115 | 2.8E-7 |
| SUPERFAMILY | SSF54368 | Glutamine synthetase, N-terminal domain | IPR036651 | Glutamine synthetase, N-terminal domain superfamily | 6 | 113 | 2.62E-17 |
| Pfam | PF00120 | Glutamine synthetase, catalytic domain | IPR008146 | Glutamine synthetase, catalytic domain | 115 | 446 | 2.5E-109 |
| Gene3D | G3DSA:3.30.590.10 | Glutamine synthetase/guanido kinase, catalytic domain | - | - | 117 | 449 | 2.2E-94 |
| FunFam | G3DSA:3.30.590.10:FF:000005 | Probable glutamine synthetase | - | - | 117 | 451 | 8.0E-125 |
| PANTHER | PTHR43785 | GAMMA-GLUTAMYLPUTRESCINE SYNTHETASE | - | - | 6 | 449 | 4.8E-95 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.